This algorithm is used by as.mo() and all the mo_* functions to determine the most probable match of taxonomic records based on user input.
Arguments
- x
Any user input value(s)
- n
A full taxonomic name, that exists in
microorganisms$fullname
Note
This algorithm was described in: Berends MS et al. (2022). AMR: An R Package for Working with Antimicrobial Resistance Data. Journal of Statistical Software, 104(3), 1-31; doi:10.18637/jss.v104.i03 .
Matching Score for Microorganisms
With ambiguous user input in as.mo() and all the mo_* functions, the returned results are chosen based on their matching score using mo_matching_score(). This matching score \(m\), is calculated as:
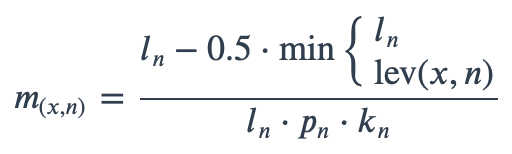
where:
x is the user input;
n is a taxonomic name (genus, species, and subspecies);
ln is the length of n;
lev is the Levenshtein distance function (counting any insertion as 1, and any deletion or substitution as 2) that is needed to change x into n;
pn is the human pathogenic prevalence group of n, as described below;
kn is the taxonomic kingdom of n, set as Bacteria = 1, Fungi = 2, Protozoa = 3, Archaea = 4, others = 5.
The grouping into human pathogenic prevalence (\(p\)) is based on experience from several microbiological laboratories in the Netherlands in conjunction with international reports on pathogen prevalence:
Group 1 (most prevalent microorganisms) consists of all microorganisms where the taxonomic class is Gammaproteobacteria or where the taxonomic genus is Enterococcus, Staphylococcus or Streptococcus. This group consequently contains all common Gram-negative bacteria, such as Pseudomonas and Legionella and all species within the order Enterobacterales.
Group 2 consists of all microorganisms where the taxonomic phylum is Proteobacteria, Firmicutes, Actinobacteria or Sarcomastigophora, or where the taxonomic genus is Absidia, Acanthamoeba, Acholeplasma, Acremonium, Actinotignum, Aedes, Alistipes, Alloprevotella, Alternaria, Amoeba, Anaerosalibacter, Ancylostoma, Angiostrongylus, Anisakis, Anopheles, Apophysomyces, Arachnia, Aspergillus, Aureobasidium, Bacteroides, Basidiobolus, Beauveria, Bergeyella, Blastocystis, Blastomyces, Borrelia, Brachyspira, Branhamella, Butyricimonas, Candida, Capillaria, Capnocytophaga, Catabacter, Cetobacterium, Chaetomium, Chlamydia, Chlamydophila, Chryseobacterium, Chrysonilia, Cladophialophora, Cladosporium, Conidiobolus, Contracaecum, Cordylobia, Cryptococcus, Curvularia, Deinococcus, Demodex, Dermatobia, Dientamoeba, Diphyllobothrium, Dirofilaria, Dysgonomonas, Echinostoma, Elizabethkingia, Empedobacter, Entamoeba, Enterobius, Exophiala, Exserohilum, Fasciola, Flavobacterium, Fonsecaea, Fusarium, Fusobacterium, Giardia, Haloarcula, Halobacterium, Halococcus, Hendersonula, Heterophyes, Histomonas, Histoplasma, Hymenolepis, Hypomyces, Hysterothylacium, Leishmania, Lelliottia, Leptosphaeria, Leptotrichia, Lucilia, Lumbricus, Malassezia, Malbranchea, Metagonimus, Meyerozyma, Microsporidium, Microsporum, Mortierella, Mucor, Mycocentrospora, Mycoplasma, Myroides, Necator, Nectria, Ochroconis, Odoribacter, Oesophagostomum, Oidiodendron, Opisthorchis, Ornithobacterium, Parabacteroides, Pediculus, Pedobacter, Phlebotomus, Phocaeicola, Phocanema, Phoma, Pichia, Piedraia, Pithomyces, Pityrosporum, Pneumocystis, Porphyromonas, Prevotella, Pseudallescheria, Pseudoterranova, Pulex, Rhizomucor, Rhizopus, Rhodotorula, Riemerella, Saccharomyces, Sarcoptes, Scolecobasidium, Scopulariopsis, Scytalidium, Sphingobacterium, Spirometra, Spiroplasma, Sporobolomyces, Stachybotrys, Streptobacillus, Strongyloides, Syngamus, Taenia, Tannerella, Tenacibaculum, Terrimonas, Toxocara, Treponema, Trichinella, Trichobilharzia, Trichoderma, Trichomonas, Trichophyton, Trichosporon, Trichostrongylus, Trichuris, Tritirachium, Trombicula, Trypanosoma, Tunga, Ureaplasma, Victivallis, Wautersiella, Weeksella or Wuchereria.
Group 3 consists of all other microorganisms.
All characters in \(x\) and \(n\) are ignored that are other than A-Z, a-z, 0-9, spaces and parentheses.
All matches are sorted descending on their matching score and for all user input values, the top match will be returned. This will lead to the effect that e.g., "E. coli" will return the microbial ID of Escherichia coli (\(m = 0.688\), a highly prevalent microorganism found in humans) and not Entamoeba coli (\(m = 0.079\), a less prevalent microorganism in humans), although the latter would alphabetically come first.
Reference Data Publicly Available
All data sets in this AMR package (about microorganisms, antibiotics, R/SI interpretation, EUCAST rules, etc.) are publicly and freely available for download in the following formats: R, MS Excel, Apache Feather, Apache Parquet, SPSS, SAS, and Stata. We also provide tab-separated plain text files that are machine-readable and suitable for input in any software program, such as laboratory information systems. Please visit our website for the download links. The actual files are of course available on our GitHub repository.
Examples
as.mo("E. coli")
#> Class 'mo'
#> [1] B_ESCHR_COLI
mo_uncertainties()
#> Matching scores are based on the resemblance between the input and the full
#> taxonomic name, and the pathogenicity in humans. See ?mo_matching_score.
#>
#> --------------------------------------------------------------------------------
#> "K. pneumoniae" -> Klebsiella pneumoniae (B_KLBSL_PNMN, 0.786)
#> Based on input "K pneumoniae"
#> Also matched: Klebsiella pneumoniae ozaenae (0.707), Klebsiella pneumoniae
#> pneumoniae (0.688), Klebsiella pneumoniae rhinoscleromatis (0.658) and
#> Kroppenstedtia pulmonis (0.304)
mo_matching_score(
x = "E. coli",
n = c("Escherichia coli", "Entamoeba coli")
)
#> [1] 0.68750000 0.07936508