Functions to plot classes sir, mic and disk, with support for base R and ggplot2.
Especially scale_x_mic() is a relevant wrapper to plot MIC values for ggplot2. It allows custom MIC ranges and to plot intermediate log2 levels for missing MIC values.
Usage
# S3 method for mic
plot(
x,
mo = NULL,
ab = NULL,
guideline = "EUCAST",
main = deparse(substitute(x)),
ylab = translate_AMR("Frequency", language = language),
xlab = translate_AMR("Minimum Inhibitory Concentration (mg/L)", language = language),
colours_SIR = c("#3CAEA3", "#F6D55C", "#ED553B"),
language = get_AMR_locale(),
expand = TRUE,
include_PKPD = getOption("AMR_include_PKPD", TRUE),
breakpoint_type = getOption("AMR_breakpoint_type", "human"),
...
)
# S3 method for mic
autoplot(
object,
mo = NULL,
ab = NULL,
guideline = "EUCAST",
title = deparse(substitute(object)),
ylab = translate_AMR("Frequency", language = language),
xlab = translate_AMR("Minimum Inhibitory Concentration (mg/L)", language = language),
colours_SIR = c("#3CAEA3", "#F6D55C", "#ED553B"),
language = get_AMR_locale(),
expand = TRUE,
include_PKPD = getOption("AMR_include_PKPD", TRUE),
breakpoint_type = getOption("AMR_breakpoint_type", "human"),
...
)
# S3 method for mic
fortify(object, ...)
scale_x_mic(
keep_operators = "edges",
mic_range = NULL,
...,
drop = FALSE,
guide = waiver(),
position = "bottom",
na.translate = TRUE
)
# S3 method for disk
plot(
x,
main = deparse(substitute(x)),
ylab = translate_AMR("Frequency", language = language),
xlab = translate_AMR("Disk diffusion diameter (mm)", language = language),
mo = NULL,
ab = NULL,
guideline = "EUCAST",
colours_SIR = c("#3CAEA3", "#F6D55C", "#ED553B"),
language = get_AMR_locale(),
expand = TRUE,
include_PKPD = getOption("AMR_include_PKPD", TRUE),
breakpoint_type = getOption("AMR_breakpoint_type", "human"),
...
)
# S3 method for disk
autoplot(
object,
mo = NULL,
ab = NULL,
title = deparse(substitute(object)),
ylab = translate_AMR("Frequency", language = language),
xlab = translate_AMR("Disk diffusion diameter (mm)", language = language),
guideline = "EUCAST",
colours_SIR = c("#3CAEA3", "#F6D55C", "#ED553B"),
language = get_AMR_locale(),
expand = TRUE,
include_PKPD = getOption("AMR_include_PKPD", TRUE),
breakpoint_type = getOption("AMR_breakpoint_type", "human"),
...
)
# S3 method for disk
fortify(object, ...)
# S3 method for sir
plot(
x,
ylab = translate_AMR("Percentage", language = language),
xlab = translate_AMR("Antimicrobial Interpretation", language = language),
main = deparse(substitute(x)),
language = get_AMR_locale(),
...
)
# S3 method for sir
autoplot(
object,
title = deparse(substitute(object)),
xlab = translate_AMR("Antimicrobial Interpretation", language = language),
ylab = translate_AMR("Frequency", language = language),
colours_SIR = c("#3CAEA3", "#F6D55C", "#ED553B"),
language = get_AMR_locale(),
...
)
# S3 method for sir
fortify(object, ...)Arguments
- x, object
values created with
as.mic(),as.disk()oras.sir()(or theirrandom_*variants, such asrandom_mic())- mo
any (vector of) text that can be coerced to a valid microorganism code with
as.mo()- ab
any (vector of) text that can be coerced to a valid antimicrobial drug code with
as.ab()- guideline
interpretation guideline to use - the default is the latest included EUCAST guideline, see Details
- main, title
title of the plot
- xlab, ylab
axis title
- colours_SIR
colours to use for filling in the bars, must be a vector of three values (in the order S, I and R). The default colours are colour-blind friendly.
- language
language to be used to translate 'Susceptible', 'Increased exposure'/'Intermediate' and 'Resistant' - the default is system language (see
get_AMR_locale()) and can be overwritten by setting the package optionAMR_locale, e.g.options(AMR_locale = "de"), see translate. Uselanguage = NULLorlanguage = ""to prevent translation.- expand
a logical to indicate whether the range on the x axis should be expanded between the lowest and highest value. For MIC values, intermediate values will be factors of 2 starting from the highest MIC value. For disk diameters, the whole diameter range will be filled.
- include_PKPD
a logical to indicate that PK/PD clinical breakpoints must be applied as a last resort - the default is
TRUE. Can also be set with the package optionAMR_include_PKPD.- breakpoint_type
the type of breakpoints to use, either "ECOFF", "animal", or "human". ECOFF stands for Epidemiological Cut-Off values. The default is
"human", which can also be set with the package optionAMR_breakpoint_type.- ...
arguments passed on to methods
- keep_operators
a character specifying how to handle operators (such as
>and<=) in the input. Accepts one of three values:"all"(orTRUE) to keep all operators,"none"(orFALSE) to remove all operators, or"edges"to keep operators only at both ends of the range.- mic_range
a manual range to plot the MIC values, e.g.,
mic_range = c(0.001, 32). UseNAto set no limit on one side, e.g.,mic_range = c(NA, 32).- drop, guide, position, na.translate
arguments passed on to
ggplot2::scale_x_discrete()
Value
The autoplot() functions return a ggplot model that is extendible with any ggplot2 function.
The fortify() functions return a data.frame as an extension for usage in the ggplot2::ggplot() function.
Details
The interpretation of "I" will be named "Increased exposure" for all EUCAST guidelines since 2019, and will be named "Intermediate" in all other cases.
For interpreting MIC values as well as disk diffusion diameters, supported guidelines to be used as input for the guideline argument are: "EUCAST 2023", "EUCAST 2022", "EUCAST 2021", "EUCAST 2020", "EUCAST 2019", "EUCAST 2018", "EUCAST 2017", "EUCAST 2016", "EUCAST 2015", "EUCAST 2014", "EUCAST 2013", "EUCAST 2012", "EUCAST 2011", "CLSI 2023", "CLSI 2022", "CLSI 2021", "CLSI 2020", "CLSI 2019", "CLSI 2018", "CLSI 2017", "CLSI 2016", "CLSI 2015", "CLSI 2014", "CLSI 2013", "CLSI 2012", and "CLSI 2011".
Simply using "CLSI" or "EUCAST" as input will automatically select the latest version of that guideline.
Examples
some_mic_values <- random_mic(size = 100)
some_disk_values <- random_disk(size = 100, mo = "Escherichia coli", ab = "cipro")
some_sir_values <- random_sir(50, prob_SIR = c(0.55, 0.05, 0.30))
plot(some_mic_values)
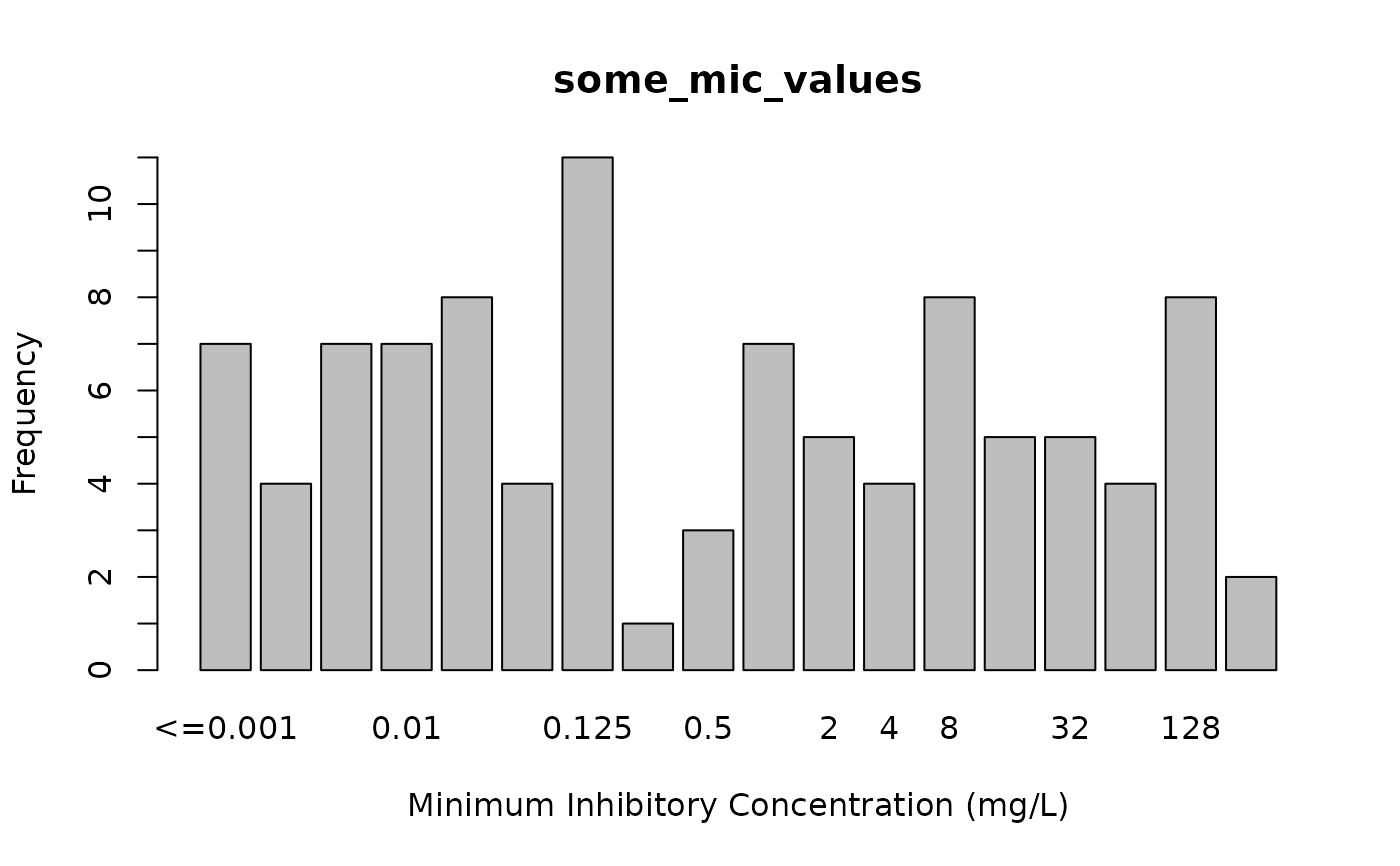 plot(some_disk_values)
plot(some_disk_values)
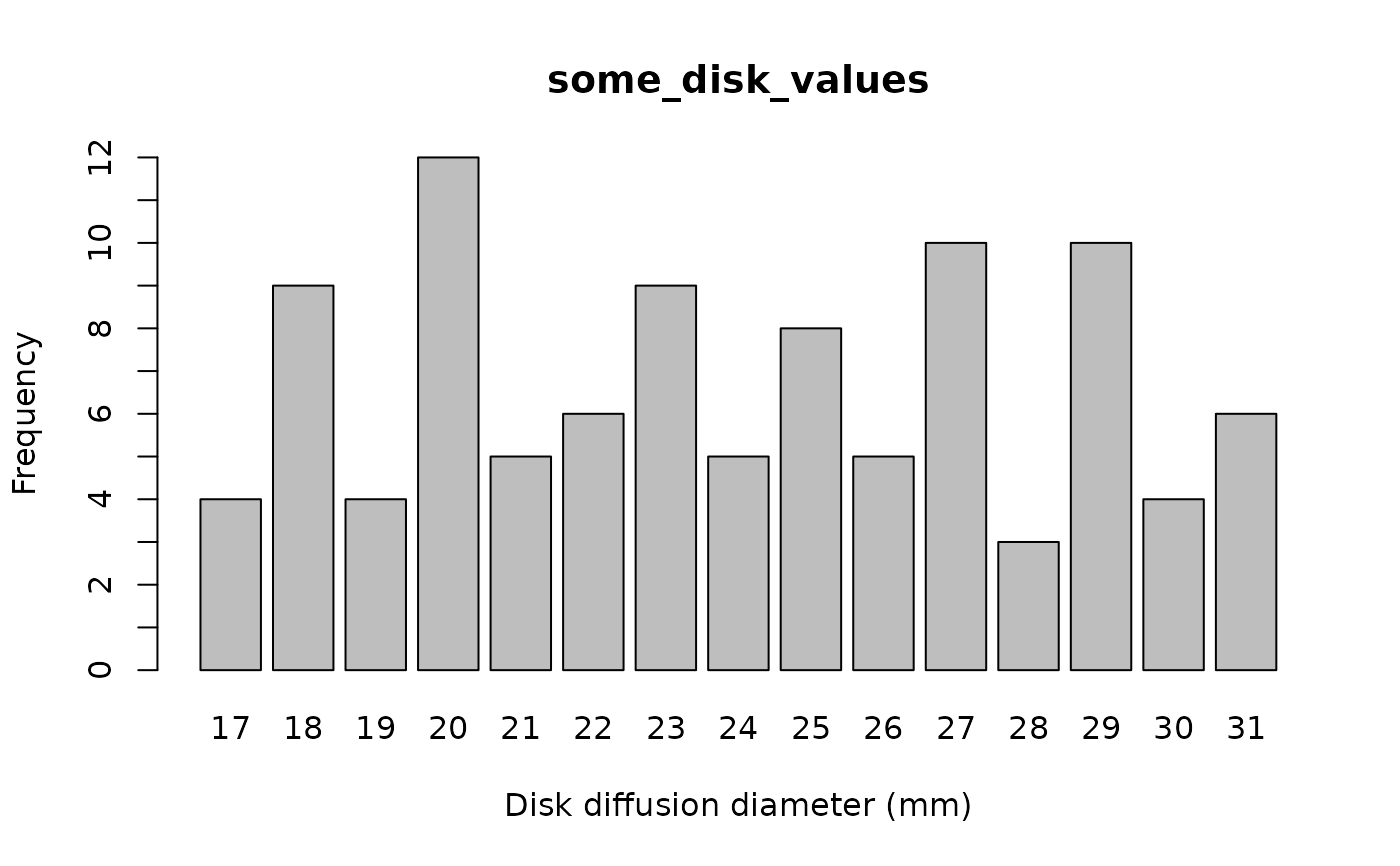 plot(some_sir_values)
plot(some_sir_values)
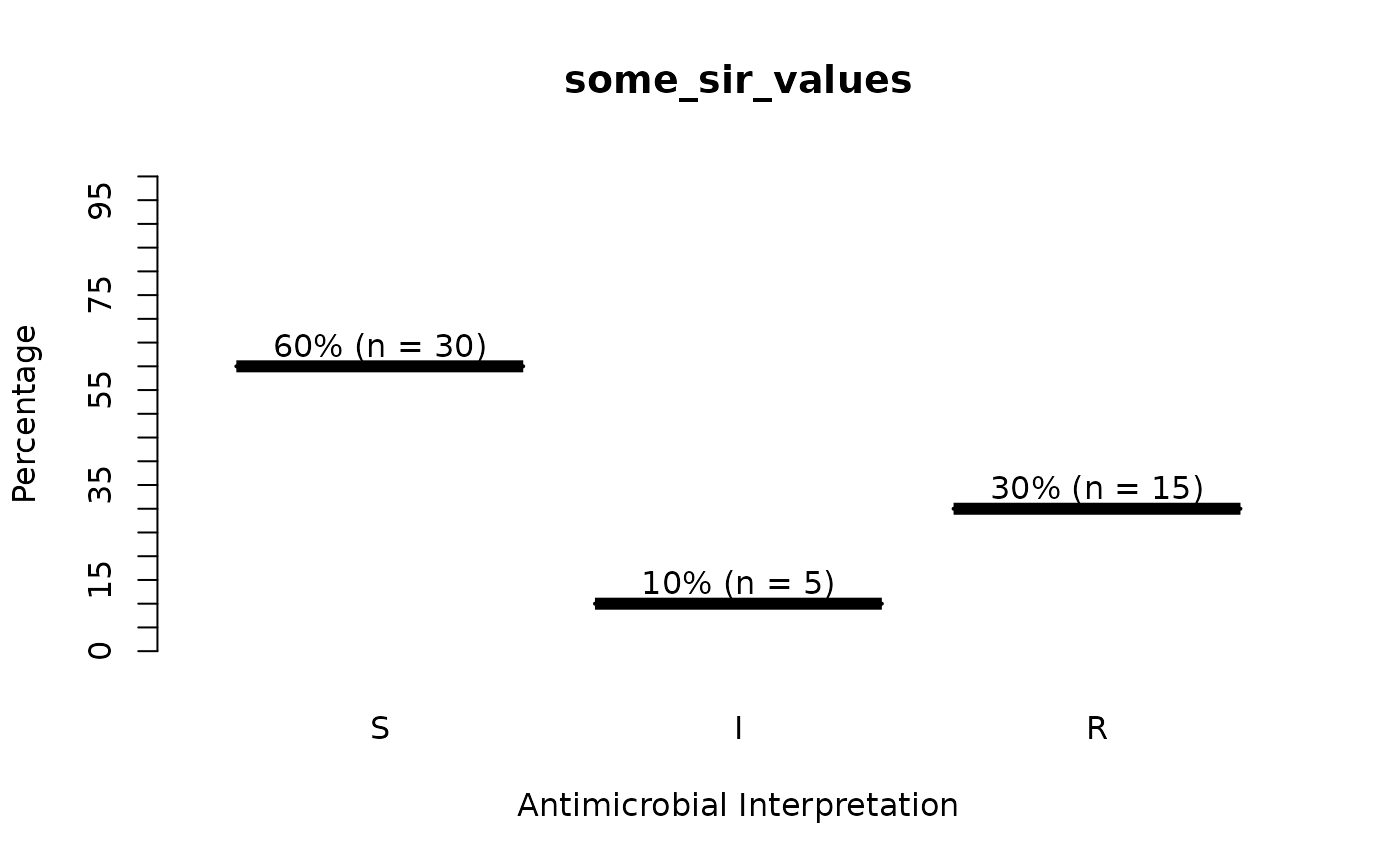 # when providing the microorganism and antibiotic, colours will show interpretations:
plot(some_mic_values, mo = "S. aureus", ab = "ampicillin")
# when providing the microorganism and antibiotic, colours will show interpretations:
plot(some_mic_values, mo = "S. aureus", ab = "ampicillin")
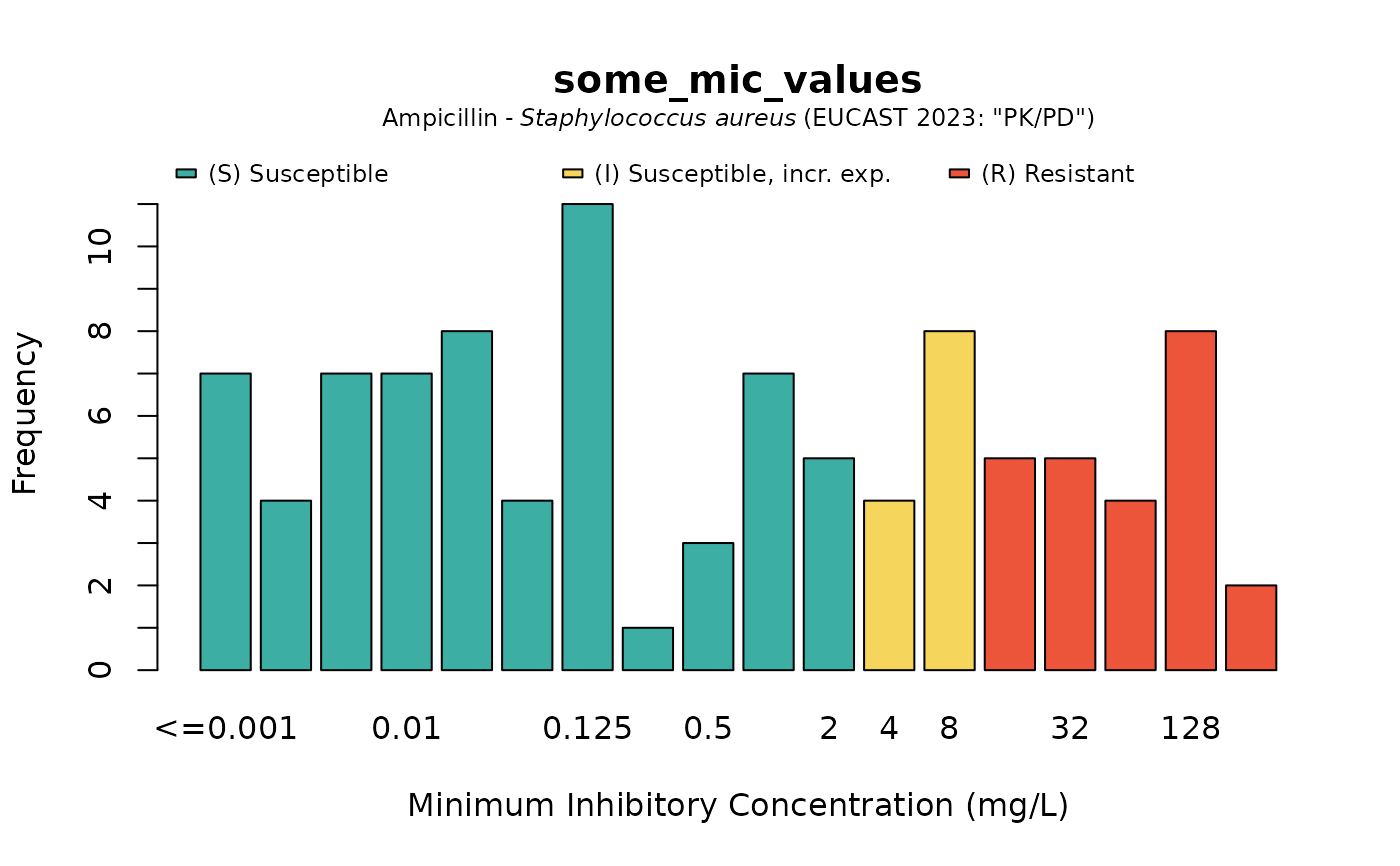 plot(some_disk_values, mo = "Escherichia coli", ab = "cipro")
plot(some_disk_values, mo = "Escherichia coli", ab = "cipro")
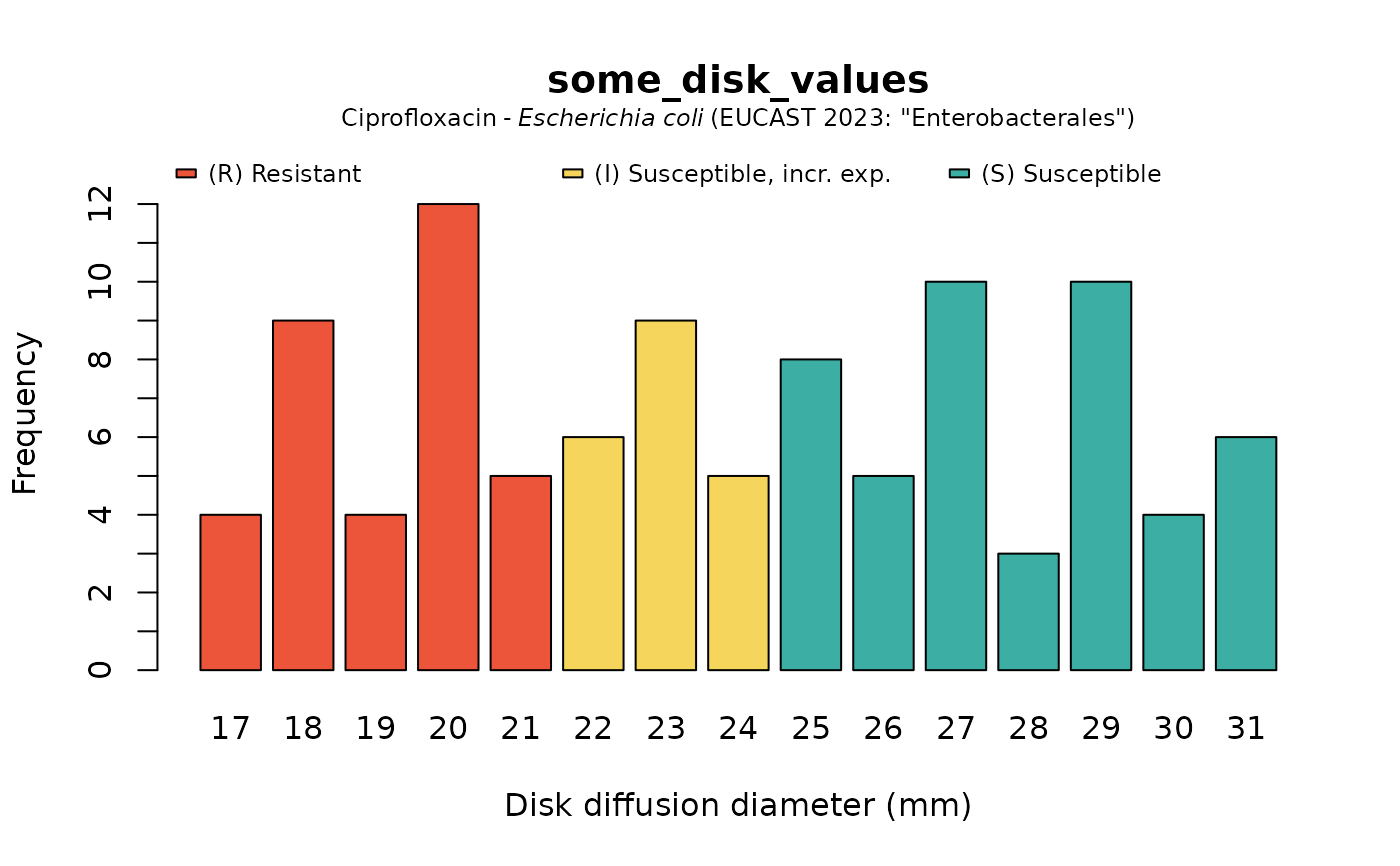 plot(some_disk_values, mo = "Escherichia coli", ab = "cipro", language = "nl")
plot(some_disk_values, mo = "Escherichia coli", ab = "cipro", language = "nl")
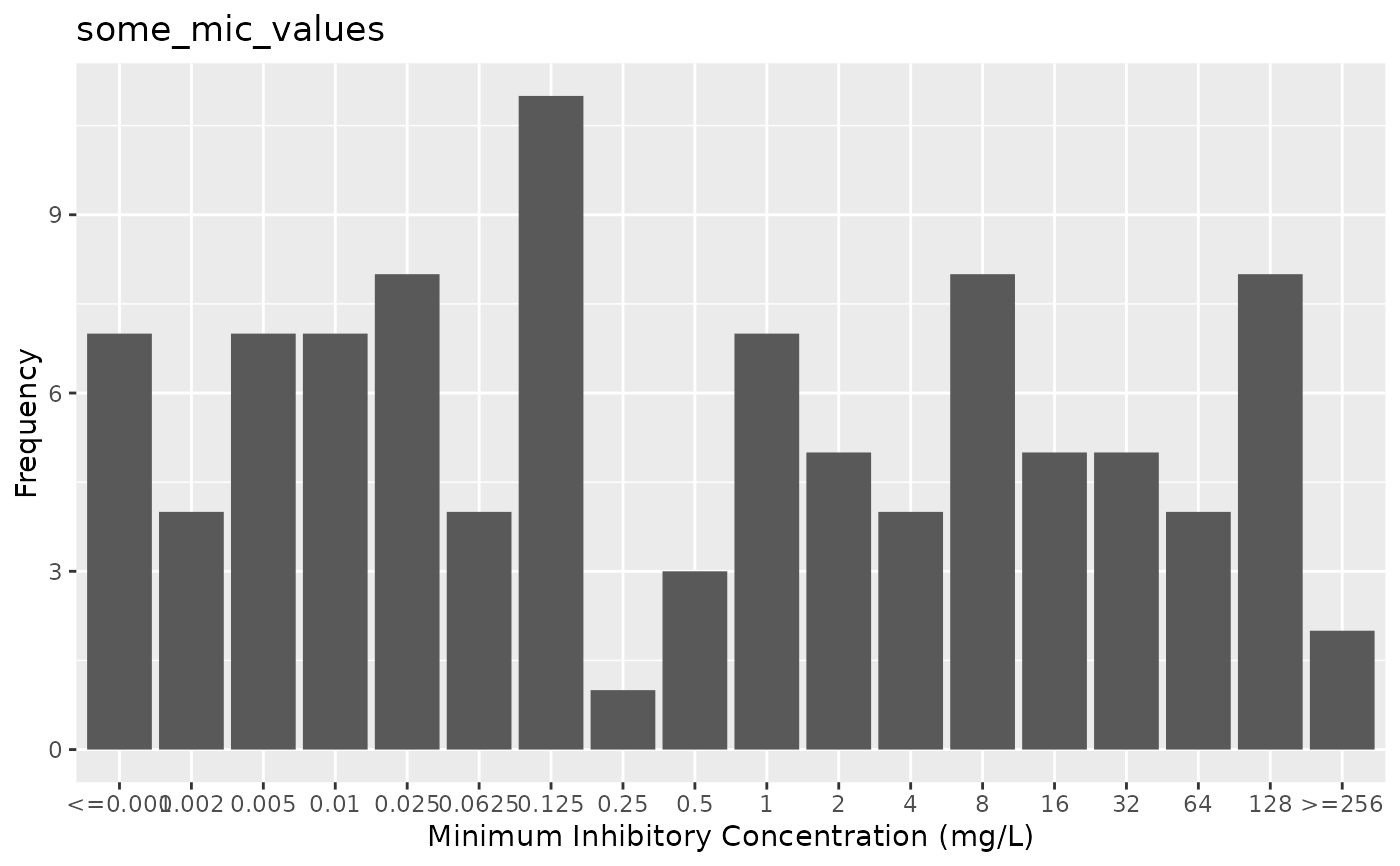
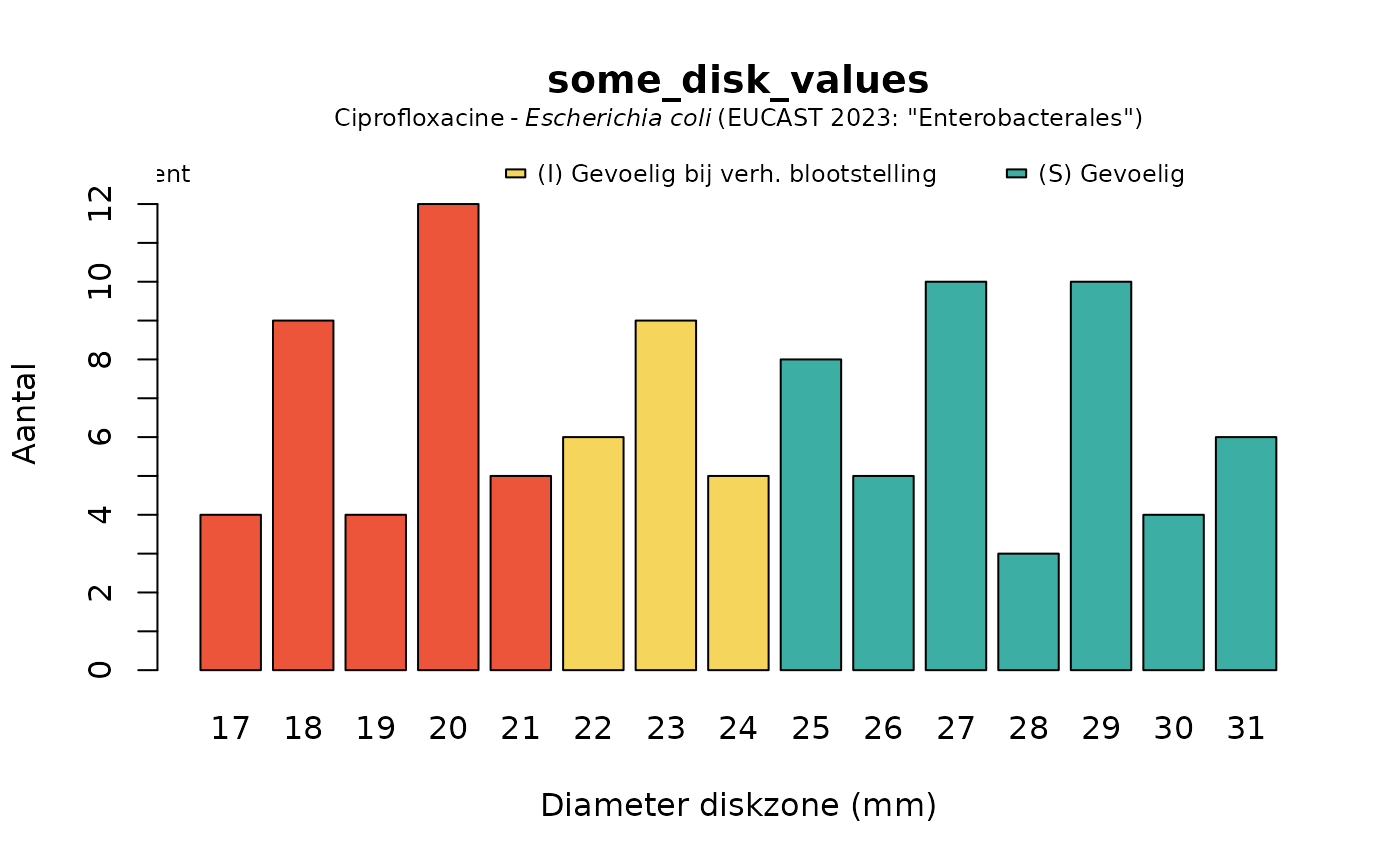 # \donttest{
if (require("ggplot2")) {
autoplot(some_mic_values)
}
# \donttest{
if (require("ggplot2")) {
autoplot(some_mic_values)
}
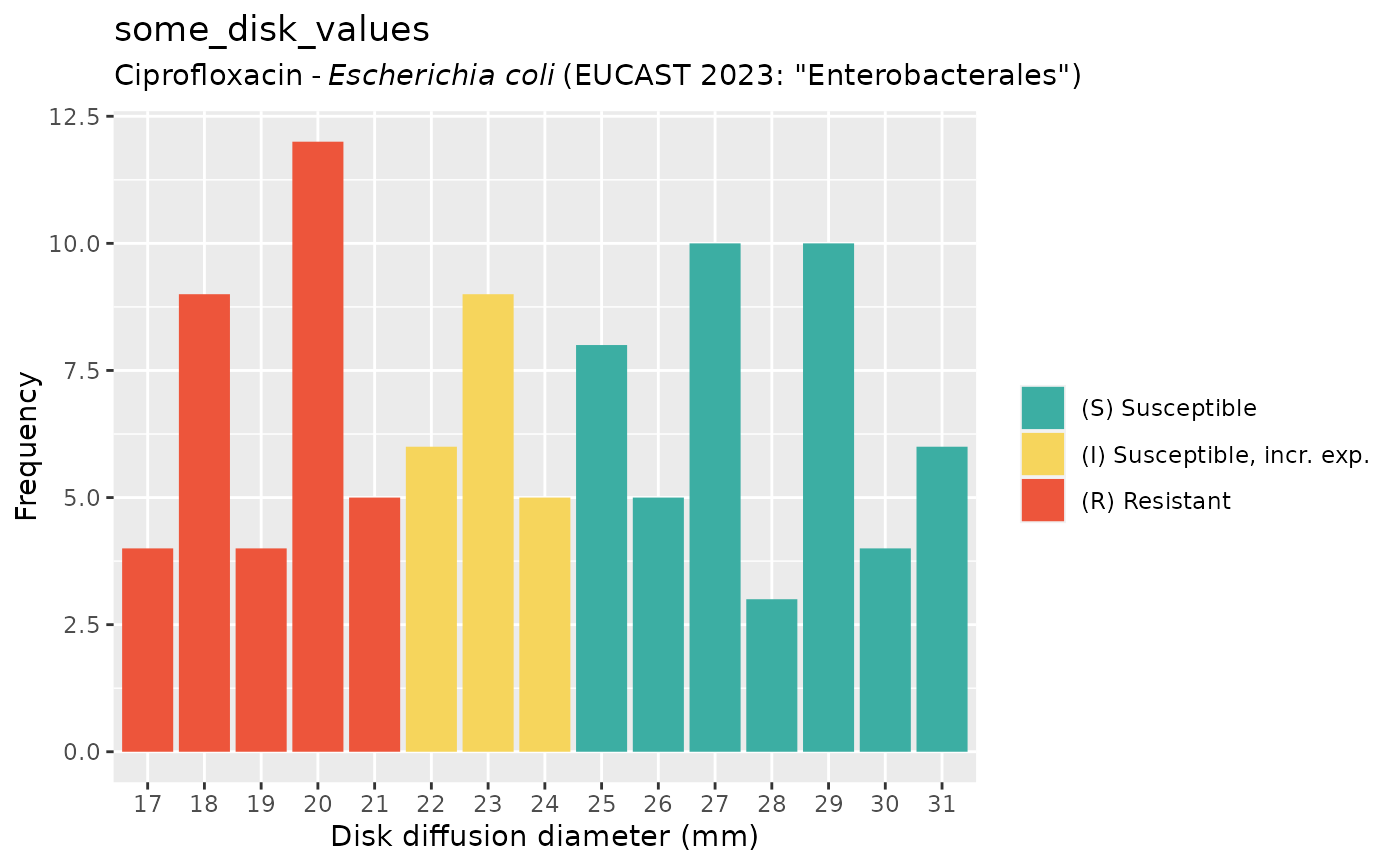
 if (require("ggplot2")) {
autoplot(some_disk_values, mo = "Escherichia coli", ab = "cipro")
}
if (require("ggplot2")) {
autoplot(some_disk_values, mo = "Escherichia coli", ab = "cipro")
}
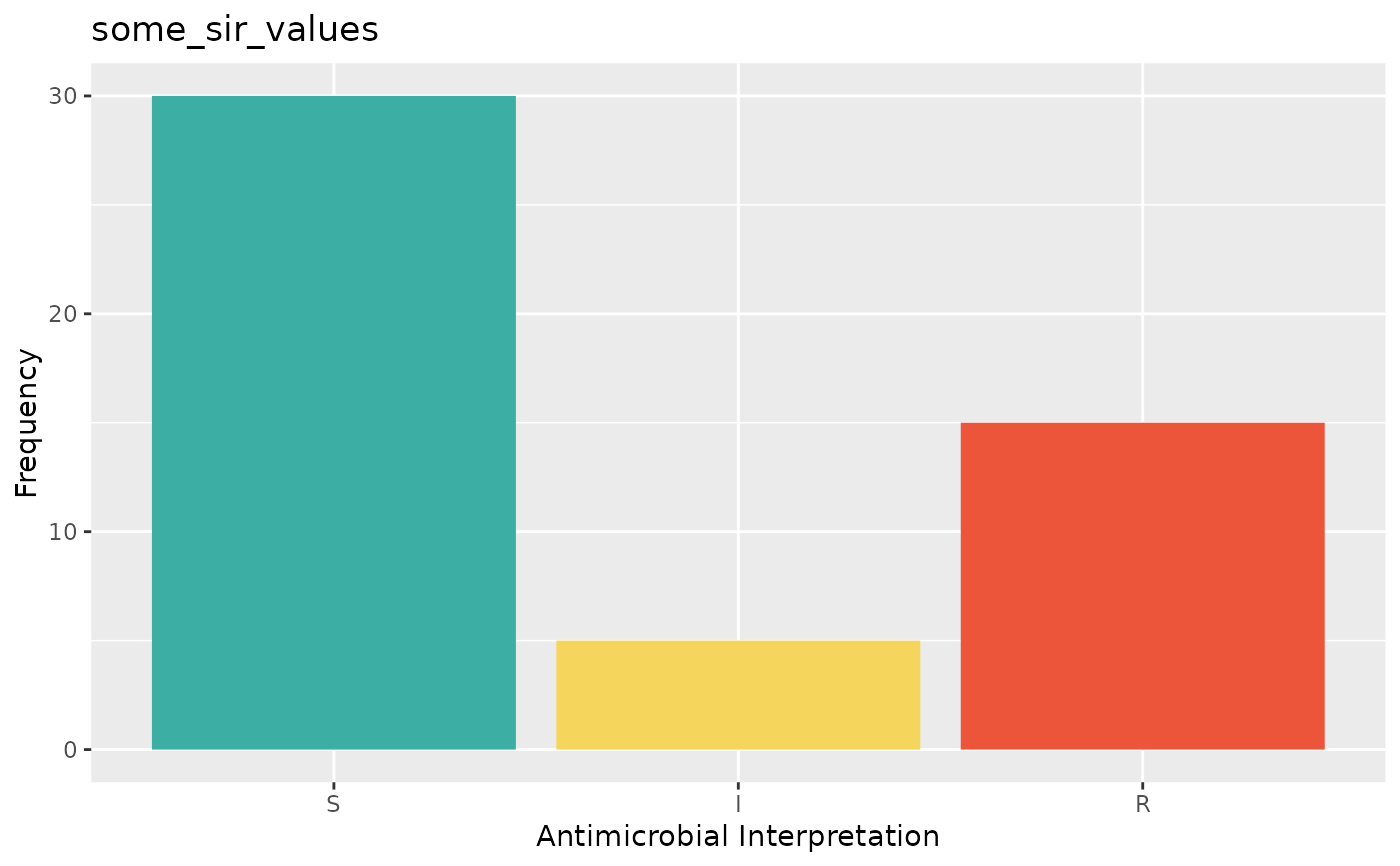
 if (require("ggplot2")) {
autoplot(some_sir_values)
}
if (require("ggplot2")) {
autoplot(some_sir_values)
}
 # }
# Plotting using scale_x_mic()
# \donttest{
if (require("ggplot2")) {
mic_plot <- ggplot(data.frame(mics = as.mic(c(0.125, "<=4", 4, 8, 32, ">=32")),
counts = c(1, 1, 2, 2, 3, 3)),
aes(mics, counts)) +
geom_col()
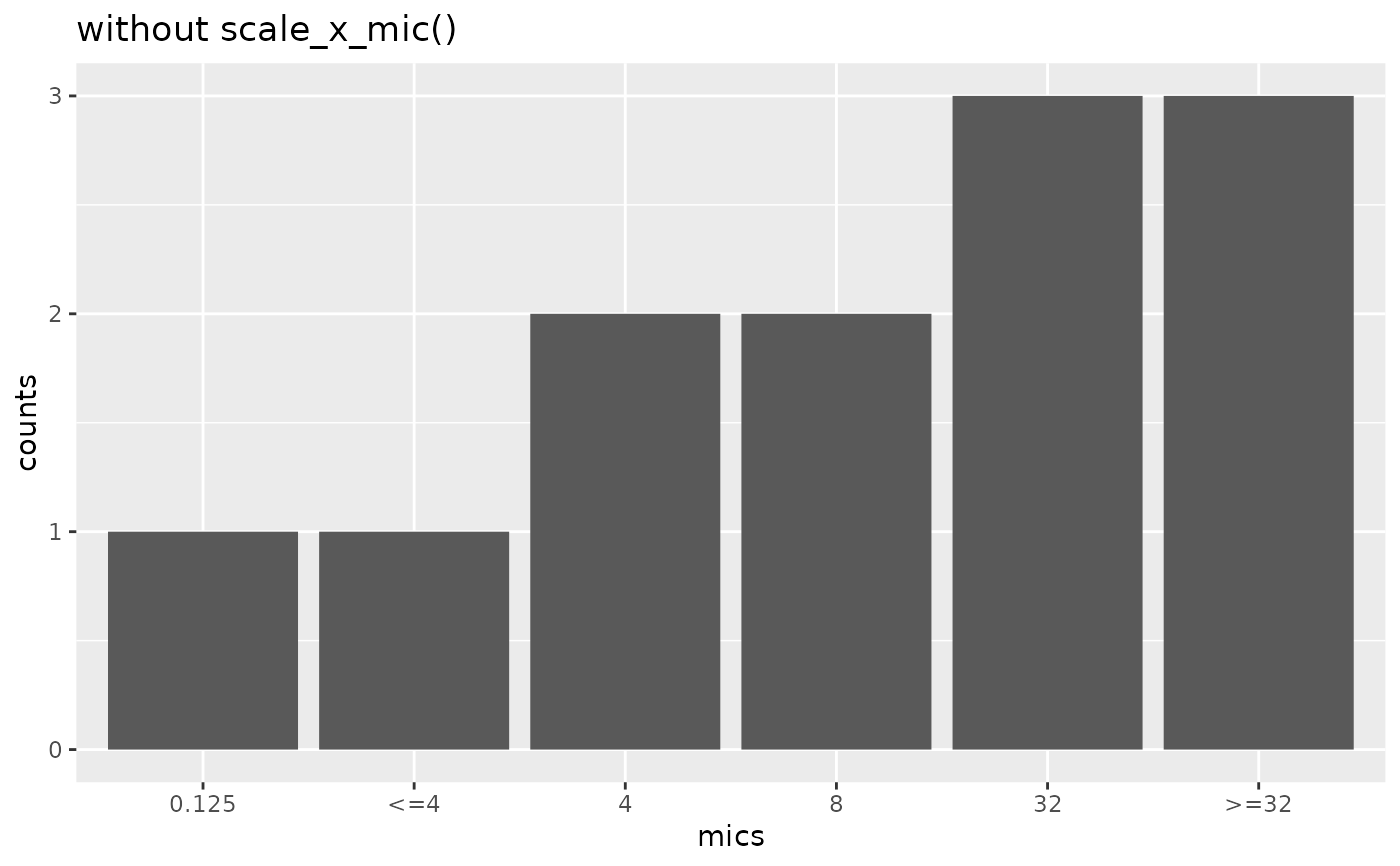
mic_plot +
labs(title = "without scale_x_mic()")
}
# }
# Plotting using scale_x_mic()
# \donttest{
if (require("ggplot2")) {
mic_plot <- ggplot(data.frame(mics = as.mic(c(0.125, "<=4", 4, 8, 32, ">=32")),
counts = c(1, 1, 2, 2, 3, 3)),
aes(mics, counts)) +
geom_col()
mic_plot +
labs(title = "without scale_x_mic()")
}
 if (require("ggplot2")) {
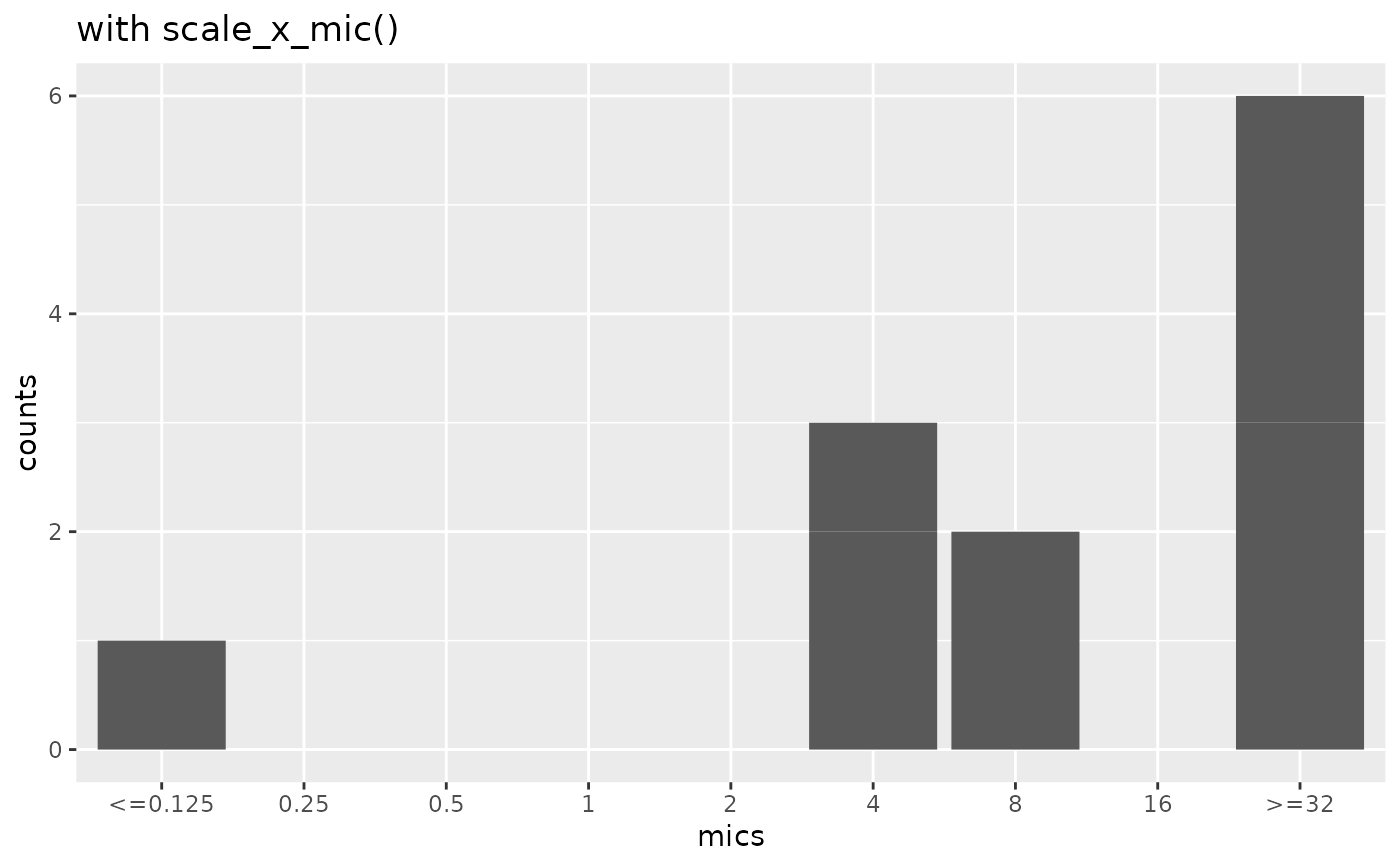
mic_plot +
scale_x_mic() +
labs(title = "with scale_x_mic()")
}
if (require("ggplot2")) {
mic_plot +
scale_x_mic() +
labs(title = "with scale_x_mic()")
}
 if (require("ggplot2")) {
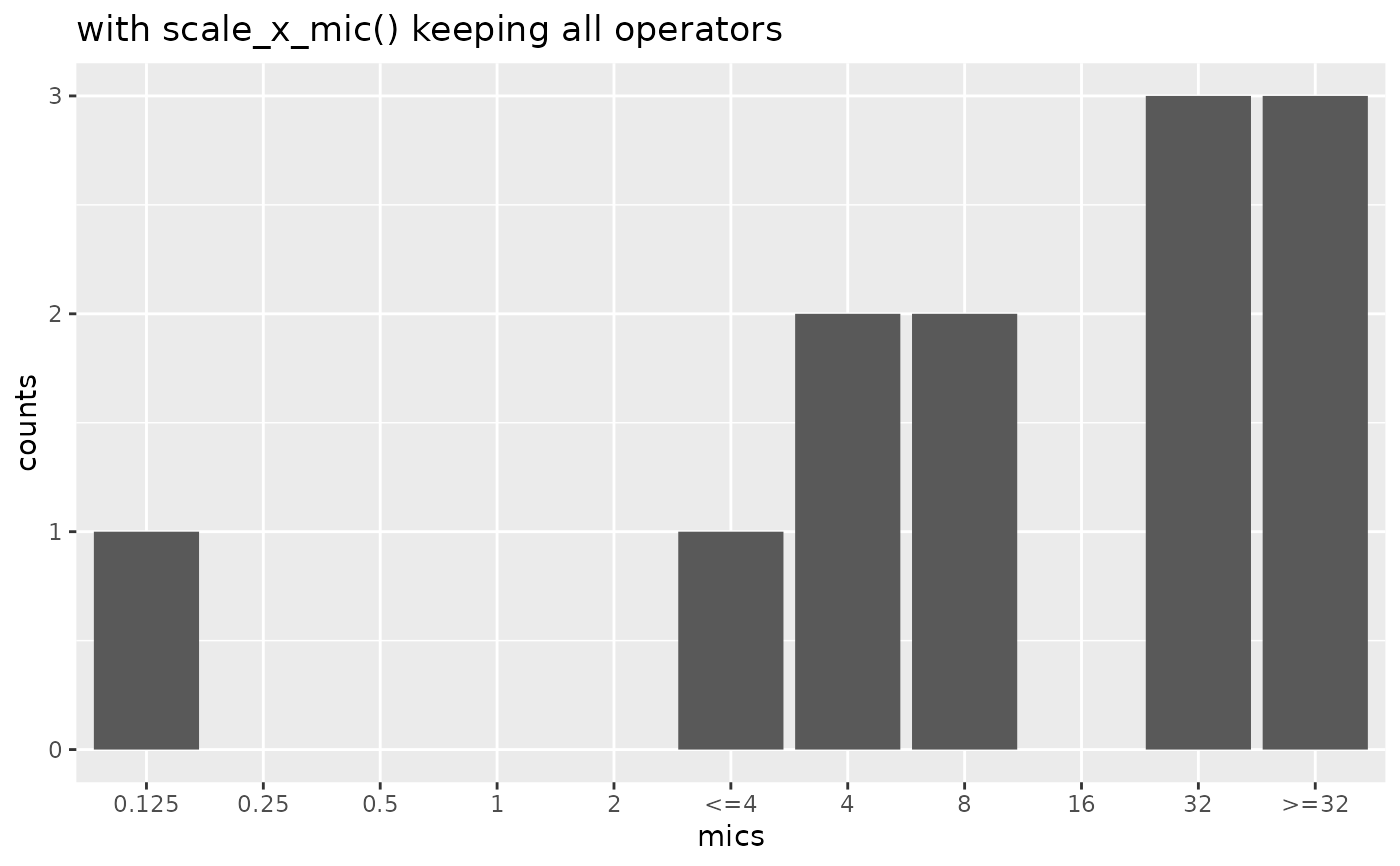
mic_plot +
scale_x_mic(keep_operators = "all") +
labs(title = "with scale_x_mic() keeping all operators")
}
if (require("ggplot2")) {
mic_plot +
scale_x_mic(keep_operators = "all") +
labs(title = "with scale_x_mic() keeping all operators")
}
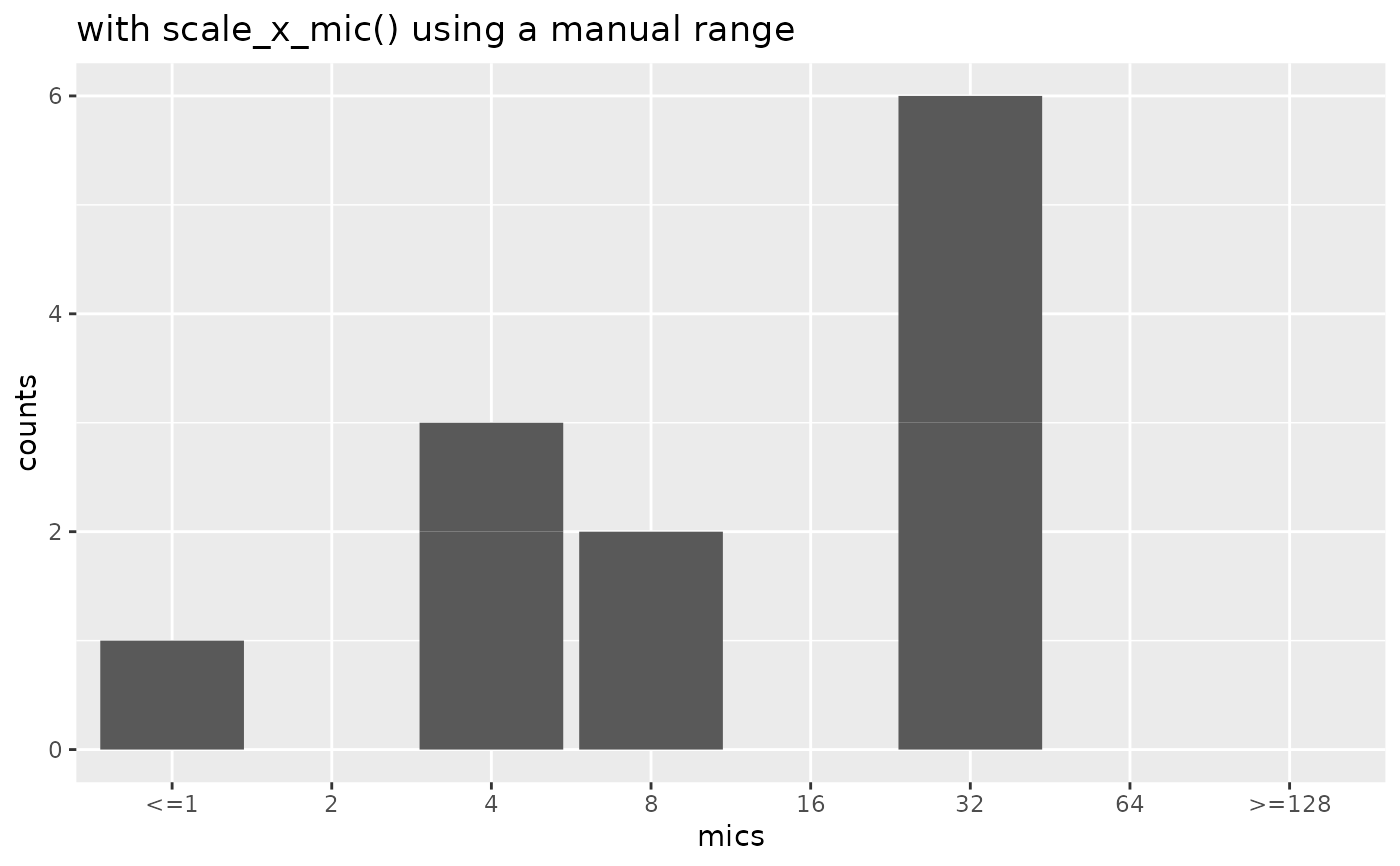
 if (require("ggplot2")) {
mic_plot +
scale_x_mic(mic_range = c(1, 128)) +
labs(title = "with scale_x_mic() using a manual range")
}
if (require("ggplot2")) {
mic_plot +
scale_x_mic(mic_range = c(1, 128)) +
labs(title = "with scale_x_mic() using a manual range")
}
 # }
# }