This transforms vectors to a new class mic, which treats the input as decimal numbers, while maintaining operators (such as ">=") and only allowing valid MIC values known to the field of (medical) microbiology.
Usage
as.mic(x, na.rm = FALSE)
NA_mic_
is.mic(x)
# S3 method for mic
droplevels(x, as.mic = FALSE, ...)Value
Ordered factor with additional class mic, that in mathematical operations acts as decimal numbers. Bare in mind that the outcome of any mathematical operation on MICs will return a numeric value.
Details
To interpret MIC values as SIR values, use as.sir() on MIC values. It supports guidelines from EUCAST (2013-2022) and CLSI (2013-2022).
This class for MIC values is a quite a special data type: formally it is an ordered factor with valid MIC values as factor levels (to make sure only valid MIC values are retained), but for any mathematical operation it acts as decimal numbers:
x <- random_mic(10)
x
#> Class 'mic'
#> [1] 16 1 8 8 64 >=128 0.0625 32 32 16
is.factor(x)
#> [1] TRUE
x[1] * 2
#> [1] 32
median(x)
#> [1] 26This makes it possible to maintain operators that often come with MIC values, such ">=" and "<=", even when filtering using numeric values in data analysis, e.g.:
x[x > 4]
#> Class 'mic'
#> [1] 16 8 8 64 >=128 32 32 16
df <- data.frame(x, hospital = "A")
subset(df, x > 4) # or with dplyr: df %>% filter(x > 4)
#> x hospital
#> 1 16 A
#> 5 64 A
#> 6 >=128 A
#> 8 32 A
#> 9 32 A
#> 10 16 AThe following generic functions are implemented for the MIC class: !, !=, %%, %/%, &, *, +, -, /, <, <=, ==, >, >=, ^, |, abs(), acos(), acosh(), all(), any(), asin(), asinh(), atan(), atanh(), ceiling(), cos(), cosh(), cospi(), cummax(), cummin(), cumprod(), cumsum(), digamma(), exp(), expm1(), floor(), gamma(), lgamma(), log(), log1p(), log2(), log10(), max(), mean(), min(), prod(), range(), round(), sign(), signif(), sin(), sinh(), sinpi(), sqrt(), sum(), tan(), tanh(), tanpi(), trigamma() and trunc(). Some functions of the stats package are also implemented: median(), quantile(), mad(), IQR(), fivenum(). Also, boxplot.stats() is supported. Since sd() and var() are non-generic functions, these could not be extended. Use mad() as an alternative, or use e.g. sd(as.numeric(x)) where x is your vector of MIC values.
Using as.double() or as.numeric() on MIC values will remove the operators and return a numeric vector. Do not use as.integer() on MIC values as by the R convention on factors, it will return the index of the factor levels (which is often useless for regular users).
Use droplevels() to drop unused levels. At default, it will return a plain factor. Use droplevels(..., as.mic = TRUE) to maintain the mic class.
NA_mic_ is a missing value of the new mic class, analogous to e.g. base R's NA_character_.
Examples
mic_data <- as.mic(c(">=32", "1.0", "1", "1.00", 8, "<=0.128", "8", "16", "16"))
mic_data
#> Class 'mic'
#> [1] >=32 1 1 1 8 <=0.128 8 16 16
is.mic(mic_data)
#> [1] TRUE
# this can also coerce combined MIC/SIR values:
as.mic("<=0.002; S")
#> Class 'mic'
#> [1] <=0.002
# mathematical processing treats MICs as numeric values
fivenum(mic_data)
#> [1] 0.128 1.000 8.000 16.000 32.000
quantile(mic_data)
#> 0% 25% 50% 75% 100%
#> 0.128 1.000 8.000 16.000 32.000
all(mic_data < 512)
#> [1] TRUE
# interpret MIC values
as.sir(
x = as.mic(2),
mo = as.mo("Streptococcus pneumoniae"),
ab = "AMX",
guideline = "EUCAST"
)
#> => Interpreting MIC values of 'AMX' (amoxicillin) according to EUCAST
#> 2022...
#> Note:
#> • Multiple breakpoints available for amoxicillin (AMX) in Streptococcus
#> pneumoniae - assuming body site 'Meningitis'.
#> Class 'sir'
#> [1] R
as.sir(
x = as.mic(c(0.01, 2, 4, 8)),
mo = as.mo("Streptococcus pneumoniae"),
ab = "AMX",
guideline = "EUCAST"
)
#> => Interpreting MIC values of 'AMX' (amoxicillin) according to EUCAST
#> 2022...
#> Note:
#> • Multiple breakpoints available for amoxicillin (AMX) in Streptococcus
#> pneumoniae - assuming body site 'Meningitis'.
#> Class 'sir'
#> [1] S R R R
# plot MIC values, see ?plot
plot(mic_data)
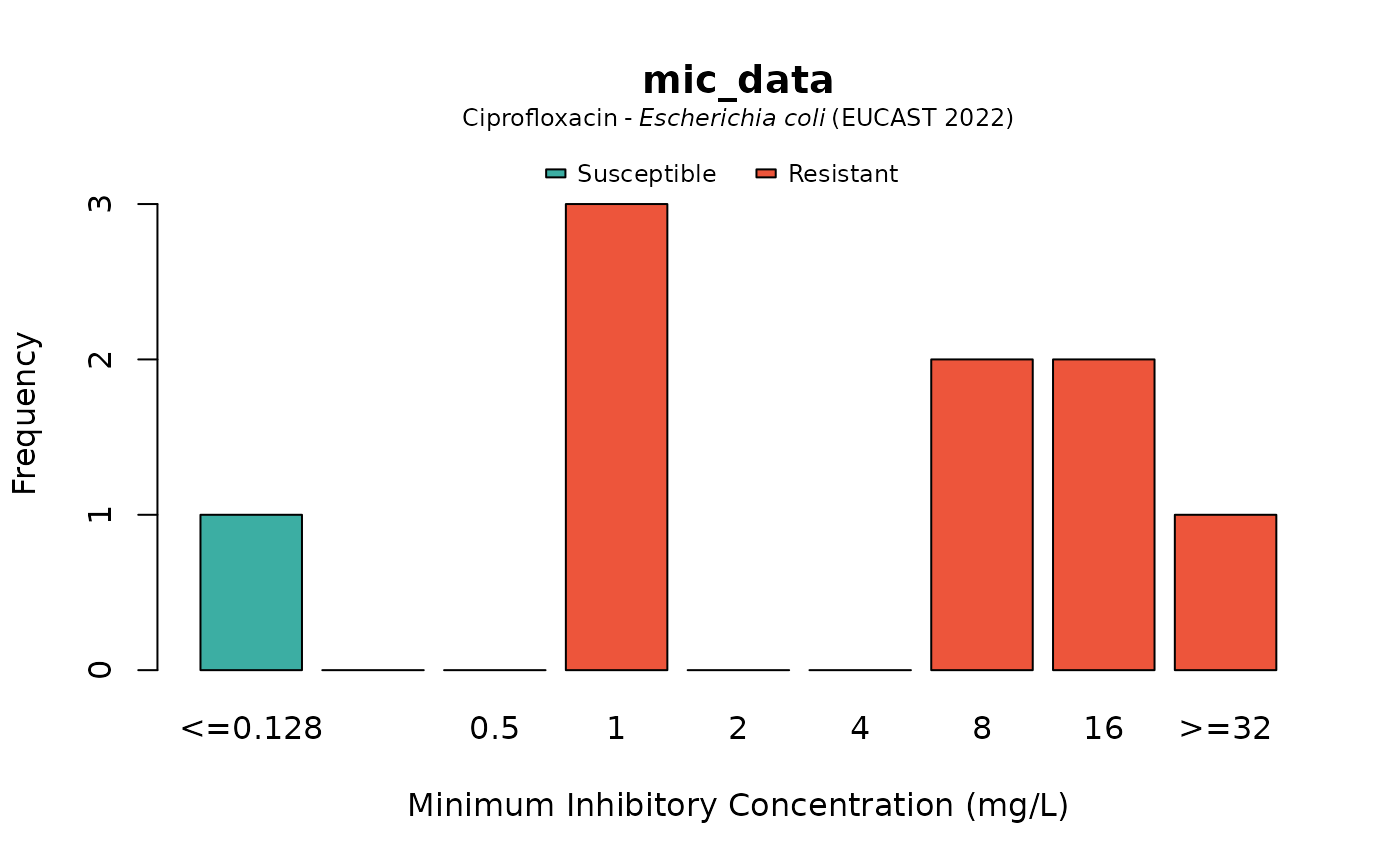
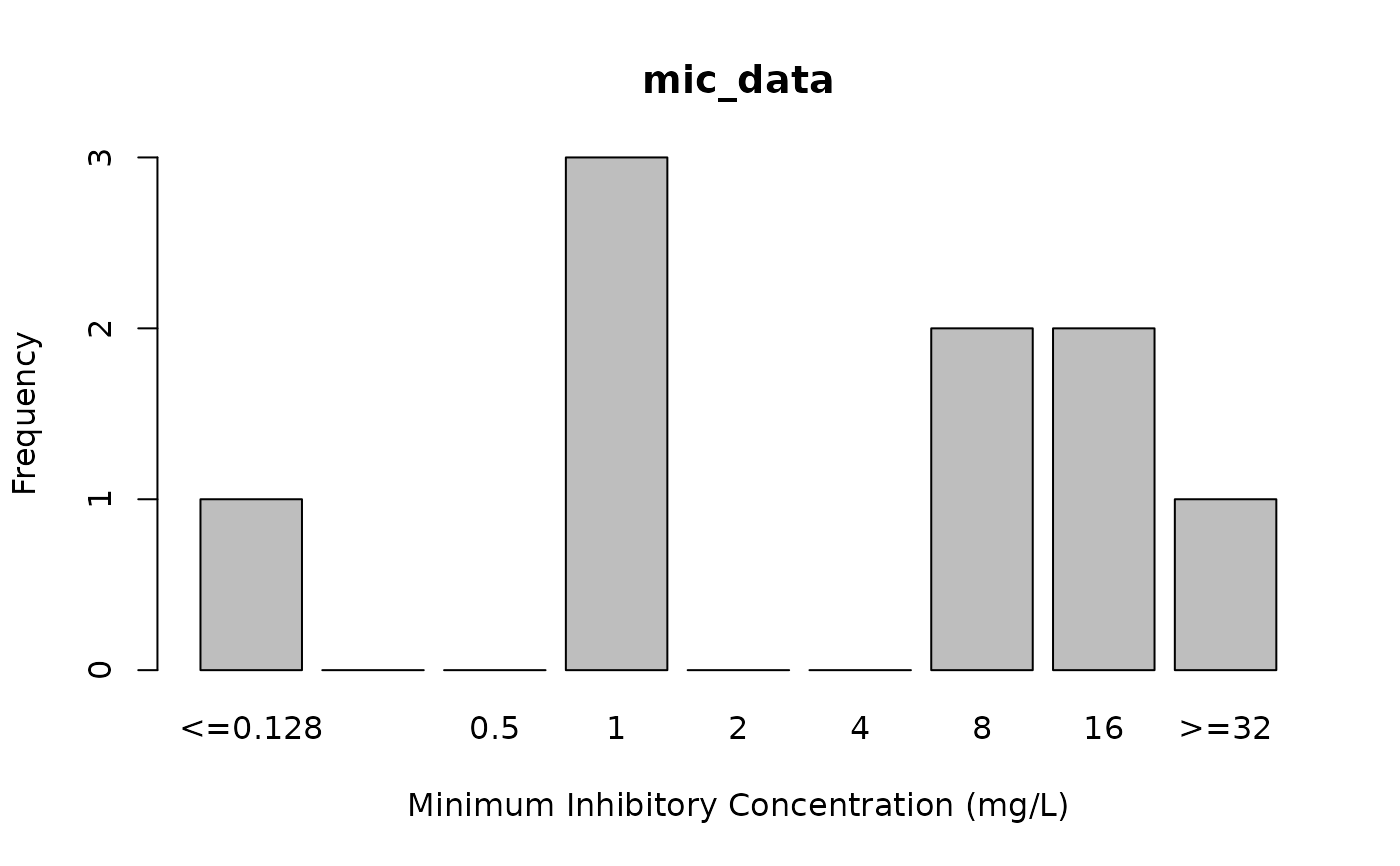 plot(mic_data, mo = "E. coli", ab = "cipro")
plot(mic_data, mo = "E. coli", ab = "cipro")
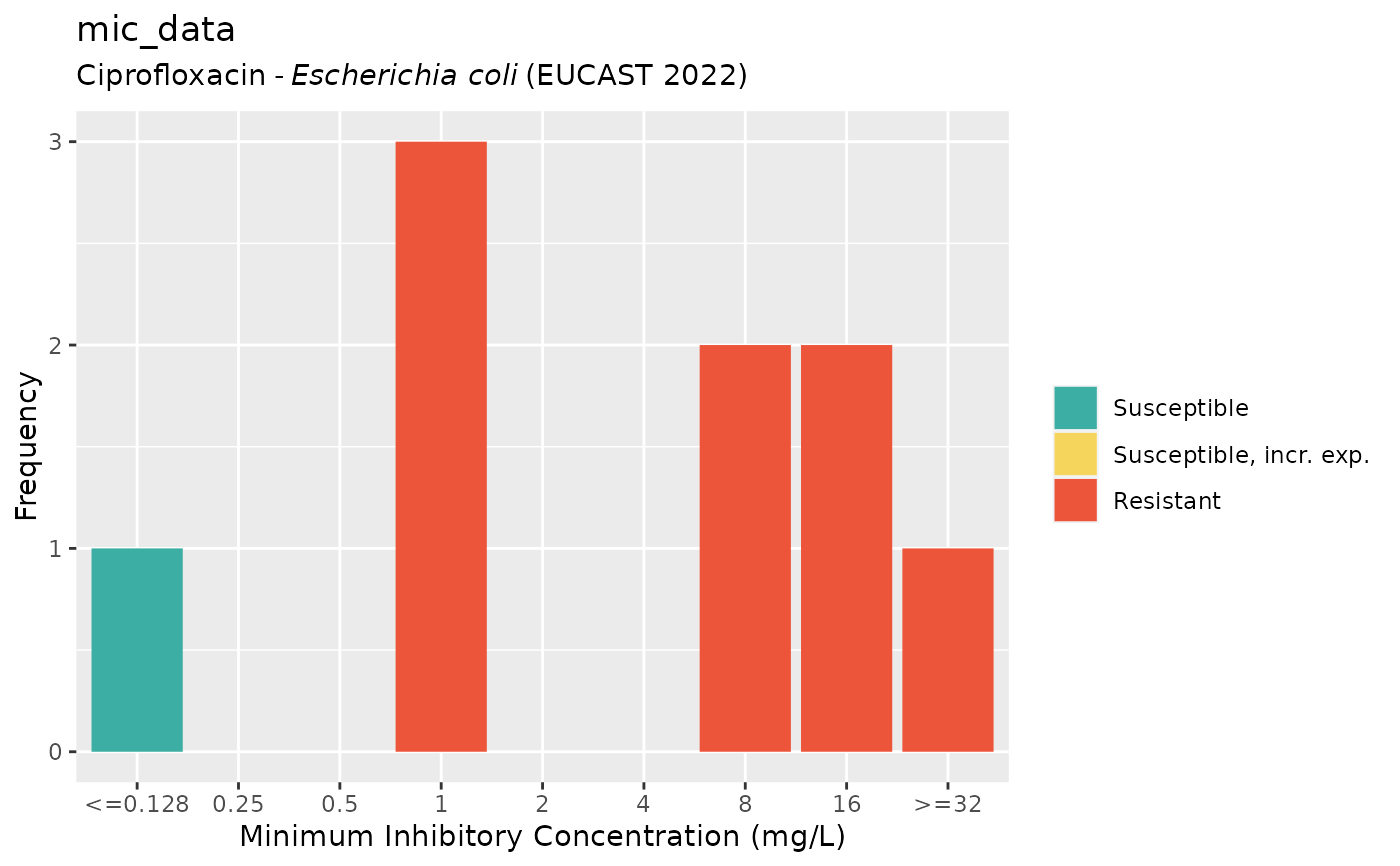
 if (require("ggplot2")) {
autoplot(mic_data, mo = "E. coli", ab = "cipro")
}
if (require("ggplot2")) {
autoplot(mic_data, mo = "E. coli", ab = "cipro")
}
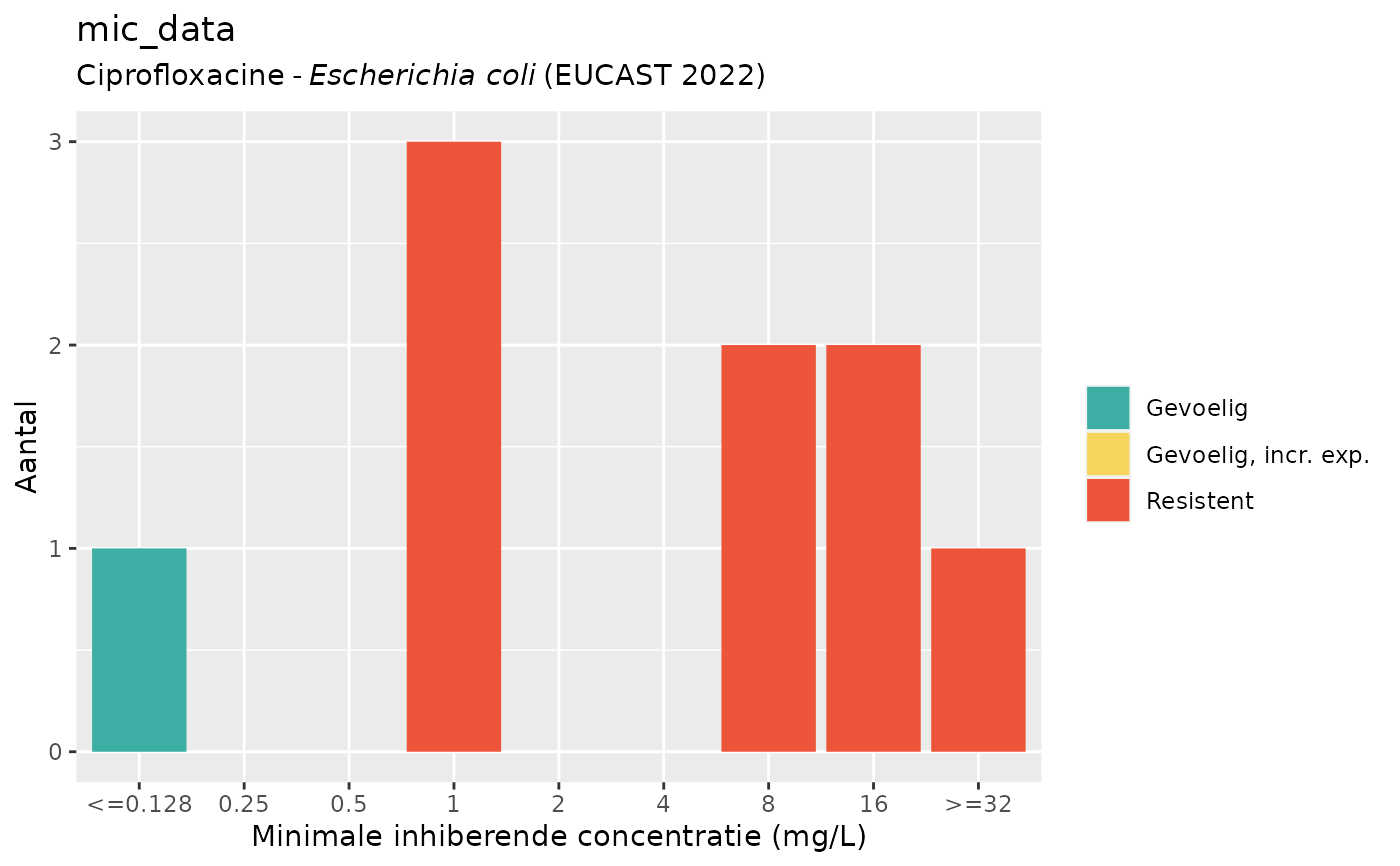
 if (require("ggplot2")) {
autoplot(mic_data, mo = "E. coli", ab = "cipro", language = "nl") # Dutch
}
if (require("ggplot2")) {
autoplot(mic_data, mo = "E. coli", ab = "cipro", language = "nl") # Dutch
}
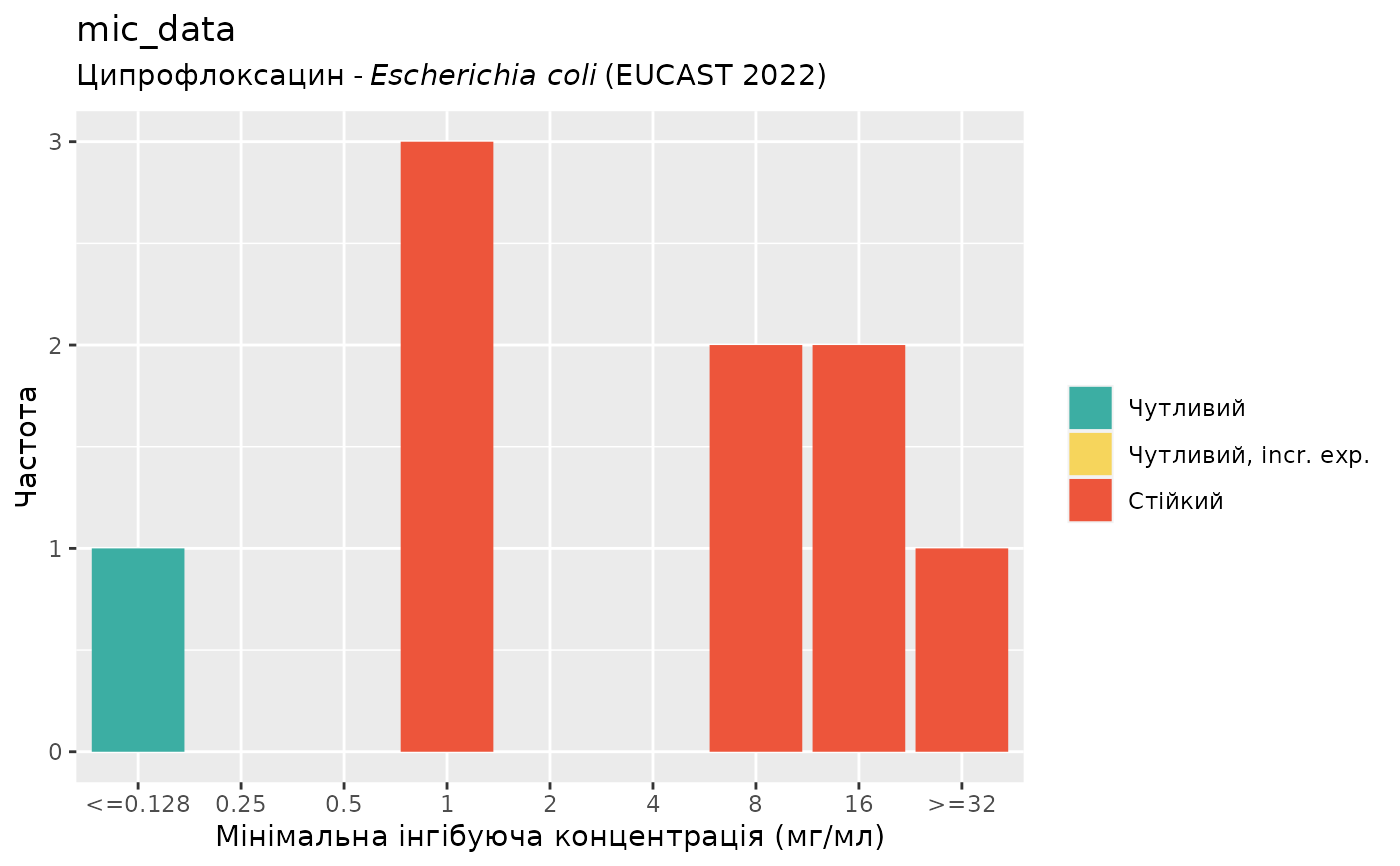
 if (require("ggplot2")) {
autoplot(mic_data, mo = "E. coli", ab = "cipro", language = "uk") # Ukrainian
}
if (require("ggplot2")) {
autoplot(mic_data, mo = "E. coli", ab = "cipro", language = "uk") # Ukrainian
}