AMR
An R package to simplify the analysis and prediction of Antimicrobial Resistance (AMR) and work with antibiotic properties by using evidence-based methods.
This R package was created for academic research by PhD students of the Faculty of Medical Sciences of the University of Groningen and the Medical Microbiology & Infection Prevention (MMBI) department of the University Medical Center Groningen (UMCG).
▶️ Download it with install.packages("AMR") or see below for other possibilities.
Authors
- Berends MS1,2, PhD Student
- Luz CF1, PhD Student
- Hassing EEA2, Data Analyst (contributor)
1 Department of Medical Microbiology, University of Groningen, University Medical Center Groningen, Groningen, the Netherlands
2 Certe Medical Diagnostics & Advice, Groningen, the Netherlands
Why this package?
This R package contains functions to make microbiological, epidemiological data analysis easier. It allows the use of some new classes to work with MIC values and antimicrobial interpretations (i.e. values S, I and R).
With AMR you can also:
- Conduct AMR analysis with the
rsifunction, that can also be used with thedplyrpackage (e.g. in conjunction withsummarise) to calculate the resistance percentages (and even co-resistance) of different antibiotic columns of a table - Predict antimicrobial resistance for the nextcoming years with the
rsi_predictfunction - Apply EUCAST rules to isolates with the
EUCAST_rulesfunction - Identify first isolates of every patient using guidelines from the CLSI (Clinical and Laboratory Standards Institute) with the
first_isolatefunction - Get antimicrobial ATC properties from the WHO Collaborating Centre for Drug Statistics Methodology (WHOCC), to be able to:
- Translate antibiotic codes (like AMOX), official names (like amoxicillin) and even trade names (like Amoxil or Trimox) to an ATC code (like J01CA04) and vice versa with the
abnamefunction - Get the latest antibiotic properties like hierarchic groups and defined daily dose (DDD) with units and administration form from the WHOCC website with the
atc_propertyfunction
- Translate antibiotic codes (like AMOX), official names (like amoxicillin) and even trade names (like Amoxil or Trimox) to an ATC code (like J01CA04) and vice versa with the
- Create frequency tables with the
freqfunction
With the MDRO function (abbreviation of Multi Drug Resistant Organisms), you can check your isolates for exceptional resistance with country-specific guidelines or EUCAST rules. Currently guidelines for Germany and the Netherlands are supported. Please suggest addition of your own country here: https://github.com/msberends/AMR/issues/new.
This package contains an example data set septic_patients, consisting of 2000 isolates from anonymised septic patients between 2001 and 2017.
How to get it?
This package is available on CRAN and also here on GitHub.
From CRAN (recommended)
-
In RStudio (recommended):
- Click on
Toolsand thenInstall Packages... - Type in
AMRand press Install
- Click on
-
install.packages("AMR")
-
In Exploratory.io:
- (Exploratory.io costs $40/month but the somewhat limited Community Plan is free for students and teachers, click here to enroll)
- Start the software and log in
- Click on your username at the right hand side top
- Click on
R Packages - Click on the
Installtab - Type in
AMRand press Install - Once it’s installed it will show up in the
User Packagessection under thePackagestab.
From GitHub (latest development version)
install.packages("devtools")
devtools::install_github("msberends/AMR")
How to use it?
# Call it with:
library(AMR)
# For a list of functions:
help(package = "AMR")
Overwrite/force resistance based on EUCAST rules
This is also called interpretive reading.
before <- data.frame(bactid = c("STAAUR", # Staphylococcus aureus
"ENCFAE", # Enterococcus faecalis
"ESCCOL", # Escherichia coli
"KLEPNE", # Klebsiella pneumoniae
"PSEAER"), # Pseudomonas aeruginosa
vanc = "-", # Vancomycin
amox = "-", # Amoxicillin
coli = "-", # Colistin
cfta = "-", # Ceftazidime
cfur = "-", # Cefuroxime
stringsAsFactors = FALSE)
before
# bactid vanc amox coli cfta cfur
# 1 STAAUR - - - - -
# 2 ENCFAE - - - - -
# 3 ESCCOL - - - - -
# 4 KLEPNE - - - - -
# 5 PSEAER - - - - -
# Now apply those rules; just need a column with bacteria ID's and antibiotic results:
after <- EUCAST_rules(before)
after
# bactid vanc amox coli cfta cfur
# 1 STAAUR - - R R -
# 2 ENCFAE - - R R R
# 3 ESCCOL R - - - -
# 4 KLEPNE R R - - -
# 5 PSEAER R R - - R
Frequency tables
Base R lacks a simple function to create frequency tables. We created such a function that works with almost all data types: freq (or frequency_tbl).
## Factors sort on item by default:
freq(septic_patients$hospital_id)
# Class: factor
# Length: 2000 (of which NA: 0 = 0.0%)
# Unique: 5
#
# Item Count Percent Cum. Count Cum. Percent (Factor Level)
# ----- ------ -------- ----------- ------------- ---------------
# A 233 11.7% 233 11.7% 1
# B 583 29.1% 816 40.8% 2
# C 221 11.1% 1037 51.8% 3
# D 650 32.5% 1687 84.4% 4
# E 313 15.7% 2000 100.0% 5
## This can be changed with the `sort.count` parameter:
freq(septic_patients$hospital_id, sort.count = TRUE)
# Class: factor
# Length: 2000 (of which NA: 0 = 0.0%)
# Unique: 5
#
# Item Count Percent Cum. Count Cum. Percent (Factor Level)
# ----- ------ -------- ----------- ------------- ---------------
# D 650 32.5% 650 32.5% 4
# B 583 29.1% 1233 61.7% 2
# E 313 15.7% 1546 77.3% 5
# A 233 11.7% 1779 88.9% 1
# C 221 11.1% 2000 100.0% 3
## Other types, like numbers or dates, sort on count by default:
> freq(septic_patients$date)
# Class: Date
# Length: 2000 (of which NA: 0 = 0.0%)
# Unique: 1662
#
# Oldest: 2 January 2001
# Newest: 18 October 2017 (+6133)
#
# Item Count Percent Cum. Count Cum. Percent
# ----------- ------ -------- ----------- -------------
# 2008-12-24 5 0.2% 5 0.2%
# 2010-12-10 4 0.2% 9 0.4%
# 2011-03-03 4 0.2% 13 0.6%
# 2013-06-24 4 0.2% 17 0.8%
# 2017-09-01 4 0.2% 21 1.1%
# 2002-09-02 3 0.2% 24 1.2%
# 2003-10-14 3 0.2% 27 1.4%
# 2004-06-25 3 0.2% 30 1.5%
# 2004-06-27 3 0.2% 33 1.7%
# 2004-10-29 3 0.2% 36 1.8%
# 2005-09-27 3 0.2% 39 2.0%
# 2006-08-01 3 0.2% 42 2.1%
# 2006-10-10 3 0.2% 45 2.2%
# 2007-11-16 3 0.2% 48 2.4%
# 2008-03-09 3 0.2% 51 2.5%
# ... and 1647 more (n = 1949; 97.5%). Use `nmax` to show more rows.
## For numeric values, some extra descriptive statistics will be calculated:
> freq(runif(n = 10, min = 1, max = 5))
# Class: numeric
# Length: 10 (of which NA: 0 = 0.0%)
# Unique: 10
#
# Mean: 3
# Std. dev.: 0.93 (CV: 0.31)
# Five-Num: 1.1 | 2.3 | 3.1 | 3.8 | 4.0 (CQV: 0.25)
# Outliers: 0
#
# Item Count Percent Cum. Count Cum. Percent
# --------- ------ -------- ----------- -------------
# 1.132033 1 10.0% 1 10.0%
# 2.226903 1 10.0% 2 20.0%
# 2.280779 1 10.0% 3 30.0%
# 2.640898 1 10.0% 4 40.0%
# 2.913462 1 10.0% 5 50.0%
# 3.364201 1 10.0% 6 60.0%
# 3.771975 1 10.0% 7 70.0%
# 3.802861 1 10.0% 8 80.0%
# 3.803547 1 10.0% 9 90.0%
# 3.985691 1 10.0% 10 100.0%
#
# Warning message:
# All observations are unique.
Learn more about this function with:
?freq
New classes
This package contains two new S3 classes: mic for MIC values (e.g. from Vitek or Phoenix) and rsi for antimicrobial drug interpretations (i.e. S, I and R). Both are actually ordered factors under the hood (an MIC of 2 being higher than <=1 but lower than >=32, and for class rsi factors are ordered as S < I < R).
Both classes have extensions for existing generic functions like print, summary and plot.
# Transform values to new classes
mic_data <- as.mic(c(">=32", "1.0", "8", "<=0.128", "8", "16", "16"))
rsi_data <- as.rsi(c(rep("S", 474), rep("I", 36), rep("R", 370)))
These functions also try to coerce valid values.
Quick overviews when just printing objects:
mic_data
# Class 'mic': 7 isolates
#
# <NA> 0
#
# <=0.128 1 8 16 >=32
# 1 1 2 2 1
rsi_data
# Class 'rsi': 880 isolates
#
# <NA>: 0
# Sum of S: 474
# Sum of IR: 406
# - Sum of R: 370
# - Sum of I: 36
#
# %S %IR %I %R
# 53.9 46.1 4.1 42.0
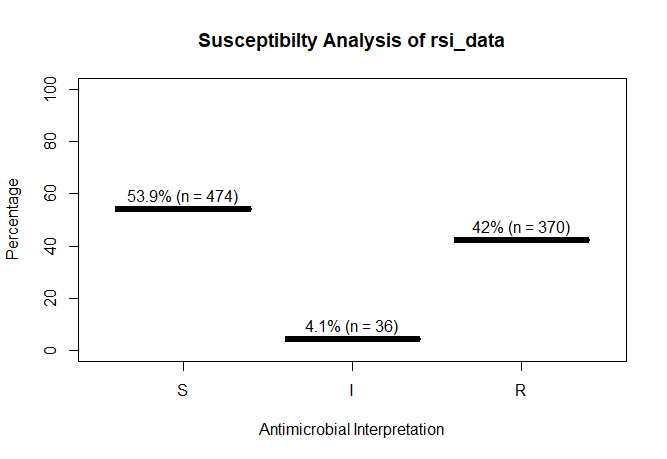
A plot of rsi_data:
plot(rsi_data)
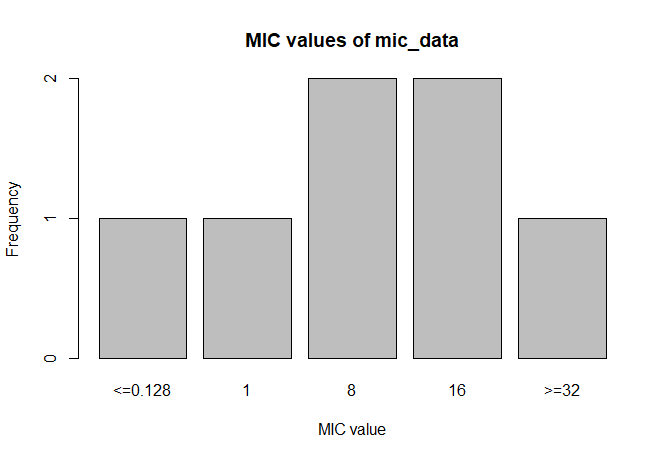
A plot of mic_data (defaults to bar plot):
plot(mic_data)
Other epidemiological functions:
# Determine key antibiotic based on bacteria ID
key_antibiotics(...)
# Selection of first isolates of any patient
first_isolate(...)
# Calculate resistance levels of antibiotics, can be used with `summarise` (dplyr)
rsi(...)
# Predict resistance levels of antibiotics
rsi_predict(...)
# Get name of antibiotic by ATC code
abname(...)
abname("J01CR02", from = "atc", to = "umcg") # "AMCL"
Databases included in package
Datasets to work with antibiotics and bacteria properties.
# Dataset with 2000 random blood culture isolates from anonymised
# septic patients between 2001 and 2017 in 5 Dutch hospitals
septic_patients # A tibble: 4,000 x 47
# Dataset with ATC antibiotics codes, official names, trade names
# and DDD's (oral and parenteral)
antibiotics # A tibble: 420 x 18
# Dataset with bacteria codes and properties like gram stain and
# aerobic/anaerobic
microorganisms # A tibble: 2,453 x 12
Copyright
This R package is licensed under the GNU General Public License (GPL) v2.0. In a nutshell, this means that this package:
-
May be used for commercial purposes
-
May be used for private purposes
-
May not be used for patent purposes
-
May be modified, although:
- Modifications must be released under the same license when distributing the package
- Changes made to the code must be documented
-
May be distributed, although:
- Source code must be made available when the package is distributed
- A copy of the license and copyright notice must be included with the package.
-
Comes with a LIMITATION of liability
-
Comes with NO warranty