Use these functions to return a specific property of a microorganism based on the latest accepted taxonomy. All input values will be evaluated internally with as.mo(), which makes it possible to use microbial abbreviations, codes and names as input. See Examples.
Usage
mo_name(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_fullname(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_shortname(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_subspecies(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_species(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_genus(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_family(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_order(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_class(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_phylum(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_kingdom(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_domain(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_type(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_status(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_gramstain(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_is_gram_negative(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_is_gram_positive(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_is_yeast(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_is_intrinsic_resistant(
x,
ab,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_snomed(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_ref(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_authors(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_year(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_lpsn(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_gbif(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_rank(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_taxonomy(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_synonyms(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_current(x, language = get_AMR_locale(), ...)
mo_info(
x,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_url(
x,
open = FALSE,
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)
mo_property(
x,
property = "fullname",
language = get_AMR_locale(),
keep_synonyms = getOption("AMR_keep_synonyms", FALSE),
...
)Arguments
- x
any character (vector) that can be coerced to a valid microorganism code with
as.mo(). Can be left blank for auto-guessing the column containing microorganism codes if used in a data set, see Examples.- language
language to translate text like "no growth", which defaults to the system language (see
get_AMR_locale())- keep_synonyms
a logical to indicate if old, previously valid taxonomic names must be preserved and not be corrected to currently accepted names. The default is
FALSE, which will return a note if old taxonomic names were processed. The default can be set withoptions(AMR_keep_synonyms = TRUE)oroptions(AMR_keep_synonyms = FALSE).- ...
other arguments passed on to
as.mo(), such as 'minimum_matching_score', 'ignore_pattern', and 'remove_from_input'- ab
any (vector of) text that can be coerced to a valid antibiotic drug code with
as.ab()- open
browse the URL using
browseURL()- property
one of the column names of the microorganisms data set: "mo", "fullname", "status", "kingdom", "phylum", "class", "order", "family", "genus", "species", "subspecies", "rank", "ref", "source", "lpsn", "lpsn_parent", "lpsn_renamed_to", "gbif", "gbif_parent", "gbif_renamed_to", "prevalence" or "snomed", or must be
"shortname"
Details
All functions will, at default, keep old taxonomic properties. Please refer to this example, knowing that Escherichia blattae was renamed to Shimwellia blattae in 2010:
mo_name("Escherichia blattae")will return"Shimwellia blattae"(with a note about the renaming)mo_ref("Escherichia blattae", keep_synonyms = TRUE)will return"Burgess et al., 1973"(without a note)mo_ref("Shimwellia blattae", keep_synonyms = FALSE)will return"Priest et al., 2010"(without a note)
The short name - mo_shortname() - almost always returns the first character of the genus and the full species, like "E. coli". Exceptions are abbreviations of staphylococci (such as "CoNS", Coagulase-Negative Staphylococci) and beta-haemolytic streptococci (such as "GBS", Group B Streptococci). Please bear in mind that e.g. E. coli could mean Escherichia coli (kingdom of Bacteria) as well as Entamoeba coli (kingdom of Protozoa). Returning to the full name will be done using as.mo() internally, giving priority to bacteria and human pathogens, i.e. "E. coli" will be considered Escherichia coli. In other words, mo_fullname(mo_shortname("Entamoeba coli")) returns "Escherichia coli".
Since the top-level of the taxonomy is sometimes referred to as 'kingdom' and sometimes as 'domain', the functions mo_kingdom() and mo_domain() return the exact same results.
Determination of the Gram stain - mo_gramstain() - will be based on the taxonomic kingdom and phylum. Originally, Cavalier-Smith defined the so-called subkingdoms Negibacteria and Posibacteria (2002, PMID 11837318), and only considered these phyla as Posibacteria: Actinobacteria, Chloroflexi, Firmicutes, and Tenericutes. These phyla were renamed to Actinomycetota, Chloroflexota, Bacillota, and Mycoplasmatota (2021, PMID 34694987). Bacteria in these phyla are considered Gram-positive in this AMR package, except for members of the class Negativicutes (within phylum Bacillota) which are Gram-negative. All other bacteria are considered Gram-negative. Species outside the kingdom of Bacteria will return a value NA. Functions mo_is_gram_negative() and mo_is_gram_positive() always return TRUE or FALSE (or NA when the input is NA or the MO code is UNKNOWN), thus always return FALSE for species outside the taxonomic kingdom of Bacteria.
Determination of yeasts - mo_is_yeast() - will be based on the taxonomic kingdom and class. Budding yeasts are fungi of the phylum Ascomycota, class Saccharomycetes (also called Hemiascomycetes). True yeasts are aggregated into the underlying order Saccharomycetales. Thus, for all microorganisms that are member of the taxonomic class Saccharomycetes, the function will return TRUE. It returns FALSE otherwise (or NA when the input is NA or the MO code is UNKNOWN).
Determination of intrinsic resistance - mo_is_intrinsic_resistant() - will be based on the intrinsic_resistant data set, which is based on 'EUCAST Expert Rules' and 'EUCAST Intrinsic Resistance and Unusual Phenotypes' v3.3 (2021). The mo_is_intrinsic_resistant() function can be vectorised over both argument x (input for microorganisms) and ab (input for antibiotics).
All output will be translated where possible.
The function mo_url() will return the direct URL to the online database entry, which also shows the scientific reference of the concerned species.
SNOMED codes - mo_snomed() - are from the version of 1 July, 2021. See Source and the microorganisms data set for more info.
Old taxonomic names (so-called 'synonyms') can be retrieved with mo_synonyms(), the current taxonomic name can be retrieved with mo_current(). Both functions return full names.
Matching Score for Microorganisms
With ambiguous user input in as.mo() and all the mo_* functions, the returned results are chosen based on their matching score using mo_matching_score(). This matching score \(m\), is calculated as:
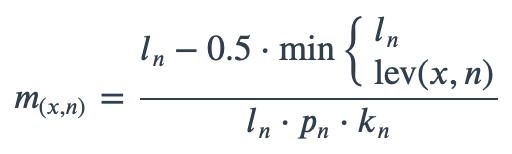
where:
x is the user input;
n is a taxonomic name (genus, species, and subspecies);
ln is the length of n;
lev is the Levenshtein distance function (counting any insertion as 1, and any deletion or substitution as 2) that is needed to change x into n;
pn is the human pathogenic prevalence group of n, as described below;
kn is the taxonomic kingdom of n, set as Bacteria = 1, Fungi = 2, Protozoa = 3, Archaea = 4, others = 5.
The grouping into human pathogenic prevalence (\(p\)) is based on recent work from Bartlett et al. (2022, doi:10.1099/mic.0.001269 ) who extensively studied medical-scientific literature to categorise all bacterial species into these groups:
Established, if a taxonomic species has infected at least three persons in three or more references. These records have
prevalence = 1.0in the microorganisms data set.Putative, if a taxonomic species has fewer than three known cases. These records have
prevalence = 2.0in the microorganisms data set.
Furthermore,
Any other bacterial genus, species or subspecies of which the genus is present in the two aforementioned groups, has
prevalence = 2.5in the microorganisms data set.Any non-bacterial genus, species or subspecies of which the genus is present in the following list, also has
prevalence = 2.5in the microorganisms data set: Absidia, Acanthamoeba, Acremonium, Aedes, Alternaria, Amoeba, Ancylostoma, Angiostrongylus, Anisakis, Anopheles, Apophysomyces, Aspergillus, Aureobasidium, Basidiobolus, Beauveria, Blastocystis, Blastomyces, Candida, Capillaria, Chaetomium, Chrysonilia, Cladophialophora, Cladosporium, Conidiobolus, Contracaecum, Cordylobia, Cryptococcus, Curvularia, Demodex, Dermatobia, Dientamoeba, Diphyllobothrium, Dirofilaria, Echinostoma, Entamoeba, Enterobius, Exophiala, Exserohilum, Fasciola, Fonsecaea, Fusarium, Giardia, Haloarcula, Halobacterium, Halococcus, Hendersonula, Heterophyes, Histomonas, Histoplasma, Hymenolepis, Hypomyces, Hysterothylacium, Leishmania, Malassezia, Malbranchea, Metagonimus, Meyerozyma, Microsporidium, Microsporum, Mortierella, Mucor, Mycocentrospora, Necator, Nectria, Ochroconis, Oesophagostomum, Oidiodendron, Opisthorchis, Pediculus, Phlebotomus, Phoma, Pichia, Piedraia, Pithomyces, Pityrosporum, Pneumocystis, Pseudallescheria, Pseudoterranova, Pulex, Rhizomucor, Rhizopus, Rhodotorula, Saccharomyces, Sarcoptes, Scolecobasidium, Scopulariopsis, Scytalidium, Spirometra, Sporobolomyces, Stachybotrys, Strongyloides, Syngamus, Taenia, Toxocara, Trichinella, Trichobilharzia, Trichoderma, Trichomonas, Trichophyton, Trichosporon, Trichostrongylus, Trichuris, Tritirachium, Trombicula, Trypanosoma, Tunga or Wuchereria.All other records have
prevalence = 3.0in the microorganisms data set.
When calculating the matching score, all characters in \(x\) and \(n\) are ignored that are other than A-Z, a-z, 0-9, spaces and parentheses.
All matches are sorted descending on their matching score and for all user input values, the top match will be returned. This will lead to the effect that e.g., "E. coli" will return the microbial ID of Escherichia coli (\(m = 0.688\), a highly prevalent microorganism found in humans) and not Entamoeba coli (\(m = 0.095\), a less prevalent microorganism in humans), although the latter would alphabetically come first.
Source
Berends MS et al. (2022). AMR: An R Package for Working with Antimicrobial Resistance Data. Journal of Statistical Software, 104(3), 1-31; doi:10.18637/jss.v104.i03
Becker K et al. (2014). Coagulase-Negative Staphylococci. Clin Microbiol Rev. 27(4): 870-926; doi:10.1128/CMR.00109-13
Becker K et al. (2019). Implications of identifying the recently defined members of the S. aureus complex, S. argenteus and S. schweitzeri: A position paper of members of the ESCMID Study Group for staphylococci and Staphylococcal Diseases (ESGS). Clin Microbiol Infect; doi:10.1016/j.cmi.2019.02.028
Becker K et al. (2020). Emergence of coagulase-negative staphylococci Expert Rev Anti Infect Ther. 18(4):349-366; doi:10.1080/14787210.2020.1730813
Lancefield RC (1933). A serological differentiation of human and other groups of hemolytic streptococci. J Exp Med. 57(4): 571-95; doi:10.1084/jem.57.4.571
Berends MS et al. (2022). Trends in Occurrence and Phenotypic Resistance of Coagulase-Negative Staphylococci (CoNS) Found in Human Blood in the Northern Netherlands between 2013 and 2019 Microorganisms 10(9), 1801; doi:10.3390/microorganisms10091801
Parte, AC et al. (2020). List of Prokaryotic names with Standing in Nomenclature (LPSN) moves to the DSMZ. International Journal of Systematic and Evolutionary Microbiology, 70, 5607-5612; doi:10.1099/ijsem.0.004332 . Accessed from https://lpsn.dsmz.de on 11 December, 2022.
GBIF Secretariat (2022). GBIF Backbone Taxonomy. Checklist dataset doi:10.15468/39omei . Accessed from https://www.gbif.org on 11 December, 2022.
Public Health Information Network Vocabulary Access and Distribution System (PHIN VADS). US Edition of SNOMED CT from 1 September 2020. Value Set Name 'Microoganism', OID 2.16.840.1.114222.4.11.1009 (v12). URL: https://phinvads.cdc.gov
Bartlett A et al. (2022). A comprehensive list of bacterial pathogens infecting humans Microbiology 168:001269; doi:10.1099/mic.0.001269
Reference Data Publicly Available
All data sets in this AMR package (about microorganisms, antibiotics, R/SI interpretation, EUCAST rules, etc.) are publicly and freely available for download in the following formats: R, MS Excel, Apache Feather, Apache Parquet, SPSS, SAS, and Stata. We also provide tab-separated plain text files that are machine-readable and suitable for input in any software program, such as laboratory information systems. Please visit our website for the download links. The actual files are of course available on our GitHub repository.
See also
Data set microorganisms
Examples
# taxonomic tree -----------------------------------------------------------
mo_kingdom("Klebsiella pneumoniae")
#> [1] "Bacteria"
mo_phylum("Klebsiella pneumoniae")
#> [1] "Pseudomonadota"
mo_class("Klebsiella pneumoniae")
#> [1] "Gammaproteobacteria"
mo_order("Klebsiella pneumoniae")
#> [1] "Enterobacterales"
mo_family("Klebsiella pneumoniae")
#> [1] "Enterobacteriaceae"
mo_genus("Klebsiella pneumoniae")
#> [1] "Klebsiella"
mo_species("Klebsiella pneumoniae")
#> [1] "pneumoniae"
mo_subspecies("Klebsiella pneumoniae")
#> [1] ""
# colloquial properties ----------------------------------------------------
mo_name("Klebsiella pneumoniae")
#> [1] "Klebsiella pneumoniae"
mo_fullname("Klebsiella pneumoniae")
#> [1] "Klebsiella pneumoniae"
mo_shortname("Klebsiella pneumoniae")
#> [1] "K. pneumoniae"
# other properties ---------------------------------------------------------
mo_gramstain("Klebsiella pneumoniae")
#> [1] "Gram-negative"
mo_snomed("Klebsiella pneumoniae")
#> [1] "1098101000112102" "1098201000112108" "409801009" "446870005"
#> [5] "56415008" "713926009" "714315002"
mo_type("Klebsiella pneumoniae")
#> [1] "Bacteria"
mo_rank("Klebsiella pneumoniae")
#> [1] "species"
mo_url("Klebsiella pneumoniae")
#> Klebsiella pneumoniae
#> "https://lpsn.dsmz.de/species/klebsiella-pneumoniae"
mo_synonyms("Klebsiella pneumoniae")
#> NULL
# scientific reference -----------------------------------------------------
mo_ref("Klebsiella pneumoniae")
#> [1] "Trevisan, 1887"
mo_authors("Klebsiella pneumoniae")
#> [1] "Trevisan"
mo_year("Klebsiella pneumoniae")
#> [1] 1887
mo_lpsn("Klebsiella pneumoniae")
#> [1] "777151"
mo_gbif("Klebsiella pneumoniae")
#> [1] "3221874"
# abbreviations known in the field -----------------------------------------
mo_genus("MRSA")
#> [1] "Staphylococcus"
mo_species("MRSA")
#> [1] "aureus"
mo_shortname("VISA")
#> [1] "S. aureus"
mo_gramstain("VISA")
#> [1] "Gram-positive"
mo_genus("EHEC")
#> [1] "Escherichia"
mo_species("EHEC")
#> [1] "coli"
# known subspecies ---------------------------------------------------------
mo_fullname("K. pneu rh")
#> [1] "Klebsiella pneumoniae rhinoscleromatis"
mo_shortname("K. pneu rh")
#> [1] "K. pneumoniae"
# \donttest{
# Becker classification, see ?as.mo ----------------------------------------
mo_fullname("Staph. epidermidis")
#> [1] "Staphylococcus epidermidis"
mo_fullname("Staph. epidermidis", Becker = TRUE)
#> [1] "Coagulase-negative Staphylococcus (CoNS)"
mo_shortname("Staph. epidermidis")
#> [1] "S. epidermidis"
mo_shortname("Staph. epidermidis", Becker = TRUE)
#> [1] "CoNS"
# Lancefield classification, see ?as.mo ------------------------------------
mo_fullname("S. pyo")
#> [1] "Streptococcus pyogenes"
mo_fullname("S. pyo", Lancefield = TRUE)
#> [1] "Streptococcus Group A"
mo_shortname("S. pyo")
#> [1] "S. pyogenes"
mo_shortname("S. pyo", Lancefield = TRUE)
#> [1] "GS. Group AS"
# language support --------------------------------------------------------
mo_gramstain("Klebsiella pneumoniae", language = "de") # German
#> [1] "Gramnegativ"
mo_gramstain("Klebsiella pneumoniae", language = "nl") # Dutch
#> [1] "Gram-negatief"
mo_gramstain("Klebsiella pneumoniae", language = "es") # Spanish
#> [1] "Gram negativo"
mo_gramstain("Klebsiella pneumoniae", language = "el") # Greek
#> [1] "Αρνητικό κατά Gram"
mo_gramstain("Klebsiella pneumoniae", language = "uk") # Ukrainian
#> [1] "Грамнегативні"
# mo_type is equal to mo_kingdom, but mo_kingdom will remain official
mo_kingdom("Klebsiella pneumoniae")
#> [1] "Bacteria"
mo_type("Klebsiella pneumoniae")
#> [1] "Bacteria"
mo_kingdom("Klebsiella pneumoniae", language = "zh") # Chinese, no effect
#> [1] "Bacteria"
mo_type("Klebsiella pneumoniae", language = "zh") # Chinese, translated
#> [1] "细菌"
mo_fullname("S. pyogenes", Lancefield = TRUE, language = "de")
#> [1] "Streptococcus Gruppe A"
mo_fullname("S. pyogenes", Lancefield = TRUE, language = "uk")
#> [1] "Streptococcus Група A"
# other --------------------------------------------------------------------
mo_is_yeast(c("Candida", "Trichophyton", "Klebsiella"))
#> [1] TRUE FALSE FALSE
# gram stains and intrinsic resistance can be used as a filter in dplyr verbs
if (require("dplyr")) {
example_isolates %>%
filter(mo_is_gram_positive()) %>%
count(mo_genus(), sort = TRUE)
}
#> ℹ Using column 'mo' as input for mo_is_gram_positive()
#> ℹ Using column 'mo' as input for mo_genus()
#> # A tibble: 18 × 2
#> `mo_genus()` n
#> <chr> <int>
#> 1 Staphylococcus 840
#> 2 Streptococcus 275
#> 3 Enterococcus 83
#> 4 Corynebacterium 17
#> 5 Micrococcus 6
#> 6 Gemella 3
#> 7 Aerococcus 2
#> 8 Cutibacterium 1
#> 9 Dermabacter 1
#> 10 Fusibacter 1
#> 11 Globicatella 1
#> 12 Granulicatella 1
#> 13 Lactobacillus 1
#> 14 Leuconostoc 1
#> 15 Listeria 1
#> 16 Paenibacillus 1
#> 17 Rothia 1
#> 18 Schaalia 1
if (require("dplyr")) {
example_isolates %>%
filter(mo_is_intrinsic_resistant(ab = "vanco")) %>%
count(mo_genus(), sort = TRUE)
}
#> ℹ Using column 'mo' as input for mo_is_intrinsic_resistant()
#> ℹ Using column 'mo' as input for mo_genus()
#> # A tibble: 20 × 2
#> `mo_genus()` n
#> <chr> <int>
#> 1 Escherichia 467
#> 2 Klebsiella 77
#> 3 Proteus 39
#> 4 Pseudomonas 30
#> 5 Serratia 25
#> 6 Enterobacter 23
#> 7 Citrobacter 11
#> 8 Haemophilus 8
#> 9 Acinetobacter 6
#> 10 Morganella 6
#> 11 Pantoea 4
#> 12 Salmonella 3
#> 13 Neisseria 2
#> 14 Stenotrophomonas 2
#> 15 Campylobacter 1
#> 16 Enterococcus 1
#> 17 Hafnia 1
#> 18 Lactobacillus 1
#> 19 Leuconostoc 1
#> 20 Pseudescherichia 1
# get a list with the complete taxonomy (from kingdom to subspecies)
mo_taxonomy("Klebsiella pneumoniae")
#> $kingdom
#> [1] "Bacteria"
#>
#> $phylum
#> [1] "Pseudomonadota"
#>
#> $class
#> [1] "Gammaproteobacteria"
#>
#> $order
#> [1] "Enterobacterales"
#>
#> $family
#> [1] "Enterobacteriaceae"
#>
#> $genus
#> [1] "Klebsiella"
#>
#> $species
#> [1] "pneumoniae"
#>
#> $subspecies
#> [1] ""
#>
# get a list with the taxonomy, the authors, Gram-stain,
# SNOMED codes, and URL to the online database
mo_info("Klebsiella pneumoniae")
#> $kingdom
#> [1] "Bacteria"
#>
#> $phylum
#> [1] "Pseudomonadota"
#>
#> $class
#> [1] "Gammaproteobacteria"
#>
#> $order
#> [1] "Enterobacterales"
#>
#> $family
#> [1] "Enterobacteriaceae"
#>
#> $genus
#> [1] "Klebsiella"
#>
#> $species
#> [1] "pneumoniae"
#>
#> $subspecies
#> [1] ""
#>
#> $status
#> [1] "accepted"
#>
#> $synonyms
#> NULL
#>
#> $gramstain
#> [1] "Gram-negative"
#>
#> $url
#> [1] "https://lpsn.dsmz.de/species/klebsiella-pneumoniae"
#>
#> $ref
#> [1] "Trevisan, 1887"
#>
#> $snomed
#> [1] "1098101000112102" "1098201000112108" "409801009" "446870005"
#> [5] "56415008" "713926009" "714315002"
#>
# }